Efficient high-resolution deletion discovery in Caenorhabditis elegans by array comparative genomic hybridization
- PMID: 17267812
- PMCID: PMC1800925
- DOI: 10.1101/gr.5690307
Efficient high-resolution deletion discovery in Caenorhabditis elegans by array comparative genomic hybridization
Abstract
We have developed array Comparative Genomic Hybridization for Caenorhabditis elegans as a means of screening for novel induced deletions in this organism. We designed three microarrays consisting of overlapping 50-mer probes to annotated exons and micro-RNAs, the first with probes to chromosomes X and II, the second with probes to chromosome II alone, and a third to the entire genome. These arrays were used to reliably detect both a large (50 kb) multigene deletion and a small (1 kb) single-gene deletion in homozygous and heterozygous samples. In one case, a deletion breakpoint was resolved to fewer than 50 bp. In an experiment designed to identify new mutations we used the X:II and II arrays to detect deletions associated with lethal mutants on chromosome II. One is an 8-kb deletion targeting the ast-1 gene on chromosome II and another is a 141-bp deletion in the gene C06A8.1. Others span large sections of the chromosome, up to >750 kb. As a further application of array Comparative Genomic Hybridization in C. elegans we used the whole-genome array to detect the extensive natural gene content variation (almost 2%) between the N2 Bristol strain and the strain CB4856, a strain isolated in Hawaii and JU258, a strain isolated in Madeira.
Figures


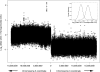




References
-
- Barstead R.J. Reverse genetics. In: Hope I.A., editor. C. elegans: A practical approach. Oxford University Press; Oxford, UK: 1999. pp. 97–118.
-
- Bartholomew G., Bailey B., Bailey B. Maui remembers: A local history. Mutual Publishing; Honolulu, HI: 1994.
-
- Chen N., Pai S., Zhao Z., Mah A., Newbury R., Johnsen R.C., Altun Z., Moerman D.G., Baillie D.L., Stein L.D., Pai S., Zhao Z., Mah A., Newbury R., Johnsen R.C., Altun Z., Moerman D.G., Baillie D.L., Stein L.D., Zhao Z., Mah A., Newbury R., Johnsen R.C., Altun Z., Moerman D.G., Baillie D.L., Stein L.D., Mah A., Newbury R., Johnsen R.C., Altun Z., Moerman D.G., Baillie D.L., Stein L.D., Newbury R., Johnsen R.C., Altun Z., Moerman D.G., Baillie D.L., Stein L.D., Johnsen R.C., Altun Z., Moerman D.G., Baillie D.L., Stein L.D., Altun Z., Moerman D.G., Baillie D.L., Stein L.D., Moerman D.G., Baillie D.L., Stein L.D., Baillie D.L., Stein L.D., Stein L.D. Identification of a nematode chemosensory gene family. Proc. Natl. Acad. Sci. 2005;102:146–151. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
