Reduced neuron-specific expression of the TAF1 gene is associated with X-linked dystonia-parkinsonism
- PMID: 17273961
- PMCID: PMC1821114
- DOI: 10.1086/512129
Reduced neuron-specific expression of the TAF1 gene is associated with X-linked dystonia-parkinsonism
Abstract
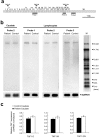
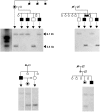
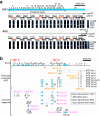
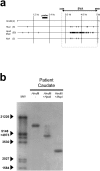
X-linked dystonia-parkinsonism (XDP) is a movement disorder endemic to the Philippines. The disease locus, DYT3, has been mapped to Xq13.1. In a search for the causative gene, we performed genomic sequencing analysis, followed by expression analysis of XDP brain tissues. We found a disease-specific SVA (short interspersed nuclear element, variable number of tandem repeats, and Alu composite) retrotransposon insertion in an intron of the TATA-binding protein-associated factor 1 gene (TAF1), which encodes the largest component of the TFIID complex, and significantly decreased expression levels of TAF1 and the dopamine receptor D2 gene (DRD2) in the caudate nucleus. We also identified an abnormal pattern of DNA methylation in the retrotransposon in the genome from the patient's caudate, which could account for decreased expression of TAF1. Our findings suggest that the reduced neuron-specific expression of the TAF1 gene is associated with XDP.
Figures



Comment in
-
The TAF1/DYT3 multiple transcript system in X-linked dystonia-parkinsonism.Am J Hum Genet. 2007 Aug;81(2):415-7; author reply 417-8. doi: 10.1086/519528. Am J Hum Genet. 2007. PMID: 17668393 Free PMC article. No abstract available.
References
Web Resources
-
- dbSNP, http://www.ncbi.nlm.nih.gov/SNP/ (for newly described SNPs ss66974122, ss66974125, ss66974128, ss66974131,ss66974134, ss66974137, ss66974140, ss66974143, ss66974146, ss66974149, ss66974152, and ss66974155)
-
- DNA Databank of Japan (DDBJ), http://www.ddbj.nig.ac.jp/Welcome-e.html (for the complete genomic sequence of the DYT3 region [accession number AB191243])
-
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/ (for XDP, Huntington disease, TAF1, ARH, FCMD, DRD2, CCFDN, DRPLA, SCA17, and TBP)
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

