Erythroid Krüppel-like factor directly activates the basic Krüppel-like factor gene in erythroid cells
- PMID: 17283065
- PMCID: PMC1899893
- DOI: 10.1128/MCB.01658-06
Erythroid Krüppel-like factor directly activates the basic Krüppel-like factor gene in erythroid cells
Abstract
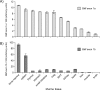
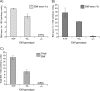
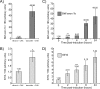
The Sp/Krüppel-like factor (Sp/Klf) family is comprised of around 25 zinc finger transcription factors that recognize CACCC boxes and GC-rich elements. We have investigated basic Krüppel-like factor (Bklf/Klf3) and show that in erythroid tissues its expression is highly dependent on another family member, erythroid Krüppel-like factor (Eklf/Klf1). We observe that Bklf mRNA is significantly reduced in erythroid tissues from Eklf-null murine embryos. We find that Bklf is driven primarily by two promoters, a ubiquitously active GC-rich upstream promoter, 1a, and an erythroid downstream promoter, 1b. Transcripts from the two promoters encode identical proteins. Interestingly, both the ubiquitous and the erythroid promoter are dependent on Eklf in erythroid cells. Eklf also activates both promoters in transient assays. Experiments utilizing an inducible form of Eklf demonstrate activation of the endogenous Bklf gene in the presence of an inhibitor of protein synthesis. The kinetics of activation are also consistent with Bklf being a direct Eklf target. Chromatin immunoprecipitation assays confirm that Eklf associates with both Bklf promoters. Eklf is typically an activator of transcription, whereas Bklf is noted as a repressor. Our results support the hypothesis that feedback cross-regulation occurs within the Sp/Klf family in vivo.
Figures





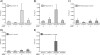
References
-
- Adam, P. J., C. P. Regan, M. B. Hautmann, and G. K. Owens. 2000. Positive- and negative-acting Kruppel-like transcription factors bind a transforming growth factor beta control element required for expression of the smooth muscle cell differentiation marker SM22alpha in vivo. J. Biol. Chem. 275:37798-37806. - PubMed
-
- Aizencang, G., C. Solis, D. F. Bishop, C. Warner, and R. J. Desnick. 2000. Human uroporphyrinogen-III synthase: genomic organization, alternative promoters, and erythroid-specific expression. Genomics 70:223-231. - PubMed
-
- Aizencang, G. I., D. F. Bishop, D. Forrest, K. H. Astrin, and R. J. Desnick. 2000. Uroporphyrinogen III synthase. An alternative promoter controls erythroid-specific expression in the murine gene. J. Biol. Chem. 275:2295-2304. - PubMed
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. - PubMed
-
- Ausubel, F. M., R. Brent, R. E. Kingston, D. D. Moore, J. G. Seidman, J. A. Smith, and K. Struhl. 1987. Current protocols in molecular biology. John Wiley and Sons, New York, NY.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
