MEK-ERK is involved in SUMO-1 foci formation on apoptosis
- PMID: 17284251
- PMCID: PMC11158486
- DOI: 10.1111/j.1349-7006.2007.00422.x
MEK-ERK is involved in SUMO-1 foci formation on apoptosis
Abstract
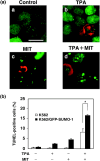
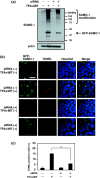
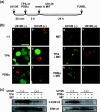
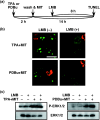
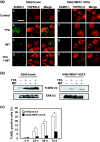
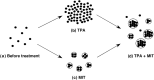
Small ubiquitin-related modifier (SUMO) modification appears to regulate the activity, intracellular localization, and stability of the targeted proteins. To explore the relationship among sumoylation, antitumor reagent, and apoptosis, we treated green fluorescence protein (GFP)-SUMO-1-overexpressed K562 cells (K562/GFP-SUMO-1) with mitoxantrone (MIT) as an antitumor reagent. By the treatment with MIT, GFP-SUMO-1 formed foci in nuclei. While by the treatment with a tumor promoter 12-O-tetradecanoylphorbol-13-acetate (TPA), GFP-SUMO-1 located homogeneously in nuclei. When K562/GFP-SUMO-1 cells were treated with TPA plus MIT, GFP-SUMO-1 foci became larger and apoptosis was induced more than with MIT alone. The apoptosis induced by TPA plus MIT was prevented by blockage of GFP-SUMO-1 foci by small interfering RNA (siRNA) against SUMO-1. The formation of GFP-SUMO-1 foci was reduced by a MEK inhibitor U0126 or a nuclear export inhibitor leptomycin B, and endogenous SUMO-1 foci were reduced in K562 cells expressing the dominant-negative MEK1 mutant. These results suggest that the formation of SUMO-1 foci is regulated by the MEK-ERK pathway and may induce apoptosis.
Figures
Similar articles
-
Growth factor-induced phosphorylation of sterol regulatory element-binding proteins inhibits sumoylation, thereby stimulating the expression of their target genes, low density lipoprotein uptake, and lipid synthesis.J Biol Chem. 2008 May 30;283(22):15224-31. doi: 10.1074/jbc.M800910200. Epub 2008 Apr 9. J Biol Chem. 2008. PMID: 18403372 Free PMC article.
-
p21WAF1 expression induced by MEK/ERK pathway activation or inhibition correlates with growth arrest, myogenic differentiation and onco-phenotype reversal in rhabdomyosarcoma cells.Mol Cancer. 2005 Dec 13;4:41. doi: 10.1186/1476-4598-4-41. Mol Cancer. 2005. PMID: 16351709 Free PMC article.
-
Requirement for ERK activation in cisplatin-induced apoptosis.J Biol Chem. 2000 Dec 15;275(50):39435-43. doi: 10.1074/jbc.M004583200. J Biol Chem. 2000. PMID: 10993883
-
Importance of MEK-1/-2 signaling in monocytic and granulocytic differentiation of myeloid cell lines.Leukemia. 2002 Apr;16(4):683-92. doi: 10.1038/sj.leu.2402400. Leukemia. 2002. PMID: 11960350
-
c-Raf/MEK/ERK pathway controls protein kinase C-mediated p70S6K activation in adult cardiac muscle cells.J Biol Chem. 2002 Jun 21;277(25):23065-75. doi: 10.1074/jbc.M200328200. Epub 2002 Apr 8. J Biol Chem. 2002. PMID: 11940578
Cited by
-
Inflammatory mediators increase SUMOylation of retinoid X receptor α in a c-Jun N-terminal kinase-dependent manner in human hepatocellular carcinoma cells.Mol Pharmacol. 2013 Aug;84(2):218-26. doi: 10.1124/mol.113.085555. Epub 2013 May 20. Mol Pharmacol. 2013. PMID: 23690070 Free PMC article.
-
In vivo analysis of a fluorescent SUMO fusion in transgenic Drosophila.Fly (Austin). 2014;8(2):108-12. doi: 10.4161/fly.28312. Fly (Austin). 2014. PMID: 25483255 Free PMC article.
References
-
- Muller S, Ledl A, Schmidt D. SUMO: a regulator of gene expression and genome integrity. Oncogene 2004; 23: 1998–2008. - PubMed
-
- Seeler JS, Dejean A. SUMO: of branched proteins and nuclear bodies. Oncogene 2001; 20: 7243–9. - PubMed
-
- Terui Y, Saad N, Jia S, McKeon F, Yuan J. Dual role of sumoylation in the nuclear localization and transcriptional activation of NFAT. J Biol Chem 2004; 279: 28 257–65. - PubMed
-
- Zhong S, Muller S, Ronchetti S, Freemont PS, Dejean A, Pandolfi PP. Role of SUMO‐1‐modified PML in nuclear body formation. Blood 2000; 95: 2748–52. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

