Increasing NADH oxidation reduces overflow metabolism in Saccharomyces cerevisiae
- PMID: 17287356
- PMCID: PMC1892921
- DOI: 10.1073/pnas.0607469104
Increasing NADH oxidation reduces overflow metabolism in Saccharomyces cerevisiae
Abstract
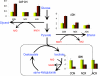
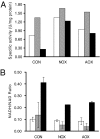
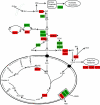
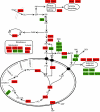
Respiratory metabolism plays an important role in energy production in the form of ATP in all aerobically growing cells. However, a limitation in respiratory capacity results in overflow metabolism, leading to the formation of byproducts, a phenomenon known as "overflow metabolism" or "the Crabtree effect." The yeast Saccharomyces cerevisiae has served as an important model organism for studying the Crabtree effect. When subjected to increasing glycolytic fluxes under aerobic conditions, there is a threshold value of the glucose uptake rate at which the metabolism shifts from purely respiratory to mixed respiratory and fermentative. It is well known that glucose repression of respiratory pathways occurs at high glycolytic fluxes, resulting in a decrease in respiratory capacity. Despite many years of detailed studies on this subject, it is not known whether the onset of the Crabtree effect is due to limited respiratory capacity or is caused by glucose-mediated repression of respiration. When respiration in S. cerevisiae was increased by introducing a heterologous alternative oxidase, we observed reduced aerobic ethanol formation. In contrast, increasing nonrespiratory NADH oxidation by overexpression of a water-forming NADH oxidase reduced aerobic glycerol formation. The metabolic response to elevated alternative oxidase occurred predominantly in the mitochondria, whereas NADH oxidase affected genes that catalyze cytosolic reactions. Moreover, NADH oxidase restored the deficiency of cytosolic NADH dehydrogenases in S. cerevisiae. These results indicate that NADH oxidase localizes in the cytosol, whereas alternative oxidase is directed to the mitochondria.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Replacing Glycerol-3-Phosphate Dehydrogenase with NADH Oxidase: Effects on Glucose Fermentation and Product Formation in Saccharomyces cerevisiae.Arch Microbiol. 2024 Nov 25;207(1):3. doi: 10.1007/s00203-024-04187-x. Arch Microbiol. 2024. PMID: 39585475
-
Metabolic impact of redox cofactor perturbations in Saccharomyces cerevisiae.Metab Eng. 2009 Jul-Sep;11(4-5):253-61. doi: 10.1016/j.ymben.2009.05.001. Epub 2009 May 13. Metab Eng. 2009. PMID: 19446033
-
Cofactor engineering in Saccharomyces cerevisiae: Expression of a H2O-forming NADH oxidase and impact on redox metabolism.Metab Eng. 2006 Jul;8(4):303-14. doi: 10.1016/j.ymben.2005.12.003. Epub 2006 Feb 10. Metab Eng. 2006. PMID: 16473032
-
Stoichiometry and compartmentation of NADH metabolism in Saccharomyces cerevisiae.FEMS Microbiol Rev. 2001 Jan;25(1):15-37. doi: 10.1111/j.1574-6976.2001.tb00570.x. FEMS Microbiol Rev. 2001. PMID: 11152939 Review.
-
Organization and regulation of the cytosolic NADH metabolism in the yeast Saccharomyces cerevisiae.Mol Cell Biochem. 2004 Jan-Feb;256-257(1-2):73-81. doi: 10.1023/b:mcbi.0000009888.79484.fd. Mol Cell Biochem. 2004. PMID: 14977171 Review.
Cited by
-
Reconstruction of cytosolic fumaric acid biosynthetic pathways in Saccharomyces cerevisiae.Microb Cell Fact. 2012 Feb 15;11:24. doi: 10.1186/1475-2859-11-24. Microb Cell Fact. 2012. PMID: 22335940 Free PMC article.
-
Requirement of the type II secretion system for utilization of cellulosic substrates by Cellvibrio japonicus.Appl Environ Microbiol. 2010 Aug;76(15):5079-87. doi: 10.1128/AEM.00454-10. Epub 2010 Jun 11. Appl Environ Microbiol. 2010. PMID: 20543053 Free PMC article.
-
Shifts in growth strategies reflect tradeoffs in cellular economics.Mol Syst Biol. 2009;5:323. doi: 10.1038/msb.2009.82. Epub 2009 Nov 3. Mol Syst Biol. 2009. PMID: 19888218 Free PMC article.
-
Deletion or overexpression of mitochondrial NAD+ carriers in Saccharomyces cerevisiae alters cellular NAD and ATP contents and affects mitochondrial metabolism and the rate of glycolysis.Appl Environ Microbiol. 2011 Apr;77(7):2239-46. doi: 10.1128/AEM.01703-10. Epub 2011 Feb 18. Appl Environ Microbiol. 2011. PMID: 21335394 Free PMC article.
-
High-Efficient Production of (S)-1-[3,5-Bis(trifluoromethyl)phenyl]ethanol via Whole-Cell Catalyst in Deep-Eutectic Solvent-Containing Micro-Aerobic Medium System.Molecules. 2020 Apr 17;25(8):1855. doi: 10.3390/molecules25081855. Molecules. 2020. PMID: 32316570 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

