Evolutionary strata on the X chromosomes of the dioecious plant Silene latifolia: evidence from new sex-linked genes
- PMID: 17287532
- PMCID: PMC1855140
- DOI: 10.1534/genetics.106.070110
Evolutionary strata on the X chromosomes of the dioecious plant Silene latifolia: evidence from new sex-linked genes
Abstract
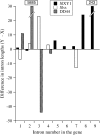
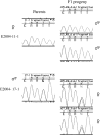
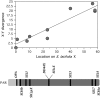
Despite its recent evolutionary origin, the sex chromosome system of the plant Silene latifolia shows signs of progressive suppression of recombination having created evolutionary strata of different X-Y divergence on sex chromosomes. However, even after 8 years of effort, this result is based on analyses of five sex-linked gene sequences, and the maximum divergence (and thus the age of this plant's sex chromosome system) has remained uncertain. More genes are therefore needed. Here, by segregation analysis of intron size variants (ISVS) and single nucleotide polymorphisms (SNPs), we identify three new Y-linked genes, one being duplicated on the Y chromosome, and test for evolutionary strata. All the new genes have homologs on the X and Y chromosomes. Synonymous divergence estimated between the X and Y homolog pairs is within the range of those already reported. Genetic mapping of the new X-linked loci shows that the map is the same in all three families that have been studied so far and that X-Y divergence increases with genetic distance from the pseudoautosomal region. We can now conclude that the divergence value is saturated, confirming the cessation of X-Y recombination in the evolution of the sex chromosomes at approximately 10-20 MYA.
Figures




Similar articles
-
Slcyt, a newly identified sex-linked gene, has recently moved onto the X chromosome in Silene latifolia (Caryophyllaceae).Mol Biol Evol. 2009 Oct;26(10):2343-51. doi: 10.1093/molbev/msp141. Epub 2009 Jul 8. Mol Biol Evol. 2009. PMID: 19587127
-
Evidence for degeneration of the Y chromosome in the dioecious plant Silene latifolia.Curr Biol. 2008 Apr 8;18(7):545-9. doi: 10.1016/j.cub.2008.03.023. Curr Biol. 2008. PMID: 18394889
-
A gradual process of recombination restriction in the evolutionary history of the sex chromosomes in dioecious plants.PLoS Biol. 2005 Jan;3(1):e4. doi: 10.1371/journal.pbio.0030004. Epub 2004 Dec 21. PLoS Biol. 2005. PMID: 15630476 Free PMC article.
-
Plant sex chromosomes: molecular structure and function.Cytogenet Genome Res. 2008;120(3-4):255-64. doi: 10.1159/000121075. Epub 2008 May 23. Cytogenet Genome Res. 2008. PMID: 18504355 Review.
-
Silene latifolia: the classical model to study heteromorphic sex chromosomes.Cytogenet Genome Res. 2010 Jul;129(1-3):250-62. doi: 10.1159/000314285. Epub 2010 Jun 10. Cytogenet Genome Res. 2010. PMID: 20551610 Review.
Cited by
-
A polymorphic pseudoautosomal boundary in the Carica papaya sex chromosomes.Mol Genet Genomics. 2015 Aug;290(4):1511-22. doi: 10.1007/s00438-015-1000-3. Epub 2015 Feb 25. Mol Genet Genomics. 2015. PMID: 25711306
-
DNA methylation and genetic degeneration of the Y chromosome in the dioecious plant Silene latifolia.BMC Genomics. 2018 Jul 16;19(1):540. doi: 10.1186/s12864-018-4936-y. BMC Genomics. 2018. PMID: 30012097 Free PMC article.
-
Evolution of sex determination and heterogamety changes in section Otites of the genus Silene.Sci Rep. 2019 Jan 31;9(1):1045. doi: 10.1038/s41598-018-37412-x. Sci Rep. 2019. PMID: 30705300 Free PMC article.
-
Molecular cytogenetic evidence of rearrangements on the Y chromosome of the threespine stickleback fish.Genetics. 2008 Aug;179(4):2173-82. doi: 10.1534/genetics.108.088559. Epub 2008 Aug 9. Genetics. 2008. PMID: 18689886 Free PMC article.
-
Expansion of the pseudo-autosomal region and ongoing recombination suppression in the Silene latifolia sex chromosomes.Genetics. 2013 Jul;194(3):673-86. doi: 10.1534/genetics.113.150755. Epub 2013 Jun 3. Genetics. 2013. PMID: 23733786 Free PMC article.
References
-
- Atanassov, I., C. Delichere, D. A. Filatov, D. Charlesworth, I. Negrutiu et al., 2001. Analysis and evolution of two functional Y-linked loci in a plant sex chromosome system. Mol. Biol. Evol. 18: 2162–2168. - PubMed
-
- Bachtrog, D., 2004. Evidence that positive selection drives Y-chromosome degeneration in Drosophila miranda. Nat. Genet. 36: 518–522. - PubMed
-
- Bradley, R. D., and D. M. Hillis, 1997. Recombinant DNA sequences generated by PCR amplification. Mol. Biol. Evol. 14: 592–593. - PubMed
-
- Carvalho, A. B., 2002. Origin and evolution of the Drosophila Y chromosome. Curr. Opin. Genet. Dev. 12: 664–668. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

