RNase MRP and the RNA processing cascade in the eukaryotic ancestor
- PMID: 17288571
- PMCID: PMC1796607
- DOI: 10.1186/1471-2148-7-S1-S13
RNase MRP and the RNA processing cascade in the eukaryotic ancestor
Abstract
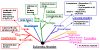
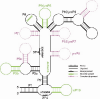
Background: Within eukaryotes there is a complex cascade of RNA-based macromolecules that process other RNA molecules, especially mRNA, tRNA and rRNA. An example is RNase MRP processing ribosomal RNA (rRNA) in ribosome biogenesis. One hypothesis is that this complexity was present early in eukaryotic evolution; an alternative is that an initial simpler network later gained complexity by gene duplication in lineages that led to animals, fungi and plants. Recently there has been a rapid increase in support for the complexity-early theory because the vast majority of these RNA-processing reactions are found throughout eukaryotes, and thus were likely to be present in the last common ancestor of living eukaryotes, herein called the Eukaryotic Ancestor.
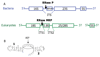
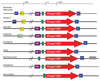
Results: We present an overview of the RNA processing cascade in the Eukaryotic Ancestor and investigate in particular, RNase MRP which was previously thought to have evolved later in eukaryotes due to its apparent limited distribution in fungi and animals and plants. Recent publications, as well as our own genomic searches, find previously unknown RNase MRP RNAs, indicating that RNase MRP has a wide distribution in eukaryotes. Combining secondary structure and promoter region analysis of RNAs for RNase MRP, along with analysis of the target substrate (rRNA), allows us to discuss this distribution in the light of eukaryotic evolution.
Conclusion: We conclude that RNase MRP can now be placed in the RNA-processing cascade of the Eukaryotic Ancestor, highlighting the complexity of RNA-processing in early eukaryotes. Promoter analyses of MRP-RNA suggest that regulation of the critical processes of rRNA cleavage can vary, showing that even these key cellular processes (for which we expect high conservation) show some species-specific variability. We present our consensus MRP-RNA secondary structure as a useful model for further searches.
Figures


Similar articles
-
Sequence analysis of RNase MRP RNA reveals its origination from eukaryotic RNase P RNA.RNA. 2006 May;12(5):699-706. doi: 10.1261/rna.2284906. Epub 2006 Mar 15. RNA. 2006. PMID: 16540690 Free PMC article.
-
Phylogenetic analysis of the structure of RNase MRP RNA in yeasts.RNA. 2002 Jun;8(6):740-51. doi: 10.1017/s1355838202022082. RNA. 2002. PMID: 12088147 Free PMC article.
-
Birth of the snoRNPs: the evolution of RNase MRP and the eukaryotic pre-rRNA-processing system.Trends Biochem Sci. 1995 Feb;20(2):78-82. doi: 10.1016/s0968-0004(00)88962-8. Trends Biochem Sci. 1995. PMID: 7701567 Review.
-
Secondary structure probing of the human RNase MRP RNA reveals the potential for MRP RNA subsets.Biochem Biophys Res Commun. 2005 Sep 23;335(2):314-21. doi: 10.1016/j.bbrc.2005.07.074. Biochem Biophys Res Commun. 2005. PMID: 16083861
-
Conserved and variable domains of RNase MRP RNA.RNA Biol. 2009 Jul-Aug;6(3):208-20. doi: 10.4161/rna.6.3.8584. Epub 2009 Jul 30. RNA Biol. 2009. PMID: 19395864 Review.
Cited by
-
Combined experimental and computational approach to identify non-protein-coding RNAs in the deep-branching eukaryote Giardia intestinalis.Nucleic Acids Res. 2007;35(14):4619-28. doi: 10.1093/nar/gkm474. Epub 2007 Jun 22. Nucleic Acids Res. 2007. PMID: 17586815 Free PMC article.
-
The emergence of predators in early life: there was no Garden of Eden.PLoS One. 2009 Jun 3;4(6):e5507. doi: 10.1371/journal.pone.0005507. PLoS One. 2009. PMID: 19492046 Free PMC article.
-
Proceedings of the First International Conference on Phylogenomics. March 15-19, 2006. Quebec, Canada.BMC Evol Biol. 2007 Feb 8;7 Suppl 1(Suppl 1):S1-16. doi: 10.1186/1471-2148-7-S1-S1. BMC Evol Biol. 2007. PMID: 17288567 Free PMC article.
-
Clustering rfam 10.1: clans, families, and classes.Genes (Basel). 2012 Jul 5;3(3):378-90. doi: 10.3390/genes3030378. Genes (Basel). 2012. PMID: 24704975 Free PMC article.
-
ptRNApred: computational identification and classification of post-transcriptional RNA.Nucleic Acids Res. 2014 Dec 16;42(22):e167. doi: 10.1093/nar/gku918. Epub 2014 Oct 10. Nucleic Acids Res. 2014. PMID: 25303994 Free PMC article.
References
-
- van Eenennaam H, Jarrous N, van Venrooij WJ, Pruijn GJ. Architecture and function of the human endonucleases RNase P and RNase MRP. IUBMB Life. 2000;49:265–272. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

