Evidence that focal adhesion complexes power bacterial gliding motility
- PMID: 17289998
- PMCID: PMC4095873
- DOI: 10.1126/science.1137223
Evidence that focal adhesion complexes power bacterial gliding motility
Abstract
The bacterium Myxococcus xanthus has two motility systems: S motility, which is powered by type IV pilus retraction, and A motility, which is powered by unknown mechanism(s). We found that A motility involved transient adhesion complexes that remained at fixed positions relative to the substratum as cells moved forward. Complexes assembled at leading cell poles and dispersed at the rear of the cells. When cells reversed direction, the A-motility clusters relocalized to the new leading poles together with S-motility proteins. The Frz chemosensory system coordinated the two motility systems. The dynamics of protein cluster localization suggest that intracellular motors and force transmission by dynamic focal adhesions can power bacterial motility.
Figures
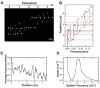



Comment in
-
Microbiology. Bright insight into bacterial gliding.Science. 2007 Feb 9;315(5813):773-4. doi: 10.1126/science.1138995. Science. 2007. PMID: 17289965 No abstract available.
References
-
- Mignot T, Merlie JP, Zusman DR. Science. 2005;310:855. - PubMed
-
-
Materials and methods are available as supporting material on Science Online
-
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

