Functional characterization of the Sindbis virus E2 glycoprotein by transposon linker-insertion mutagenesis
- PMID: 17306321
- PMCID: PMC1959473
- DOI: 10.1016/j.virol.2007.01.006
Functional characterization of the Sindbis virus E2 glycoprotein by transposon linker-insertion mutagenesis
Abstract
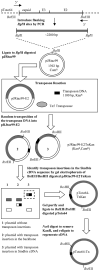
The glycoprotein envelope of alphaviruses consists of two proteins, E1 and E2. E1 is responsible for fusion and E2 is responsible for receptor binding. An atomic structure is available for E1, but one for E2 has not been reported. In this study, transposon linker-insertion mutagenesis was used to probe the function of different domains of E2. A library of mutants, containing 19 amino acid insertions in the E2 glycoprotein sequence of the prototype alphavirus, Sindbis virus (SINV), was generated. Fifty-seven independent E2 insertions were characterized, of which more than half (67%) gave rise to viable virus. The wild-type-like mutants identify regions that accommodate insertions without perturbing virus production and can be used to insert targeting moieties to direct SINV to specific receptors. The defective and lethal mutants give insight into regions of E2 important for protein stability, transport to the cell membrane, E1-E2 contacts, and receptor binding.
Figures



Similar articles
-
Mutating conserved cysteines in the alphavirus e2 glycoprotein causes virus-specific assembly defects.J Virol. 2012 Mar;86(6):3100-11. doi: 10.1128/JVI.06615-11. Epub 2012 Jan 11. J Virol. 2012. PMID: 22238319 Free PMC article.
-
Sindbis virus attachment: isolation and characterization of mutants with impaired binding to vertebrate cells.J Virol. 1993 Jun;67(6):3363-74. doi: 10.1128/JVI.67.6.3363-3374.1993. J Virol. 1993. PMID: 7684466 Free PMC article.
-
Structural rearrangement of infecting Sindbis virions at the cell surface: mapping of newly accessible epitopes.J Virol. 1993 Sep;67(9):5117-25. doi: 10.1128/JVI.67.9.5117-5125.1993. J Virol. 1993. PMID: 7688818 Free PMC article.
-
Use of a lambda gt11 expression library to localize a neutralizing antibody-binding site in glycoprotein E2 of Sindbis virus.J Virol. 1991 Dec;65(12):7037-40. doi: 10.1128/JVI.65.12.7037-7040.1991. J Virol. 1991. PMID: 1719239 Free PMC article.
-
Comprehensive linker-scanning mutagenesis of the hepatitis C virus E1 and E2 envelope glycoproteins reveals new structure-function relationships.J Gen Virol. 2011 Oct;92(Pt 10):2249-2261. doi: 10.1099/vir.0.034314-0. Epub 2011 Jun 22. J Gen Virol. 2011. PMID: 21697343 Free PMC article.
Cited by
-
Genetic determinants of Sindbis virus mosquito infection are associated with a highly conserved alphavirus and flavivirus envelope sequence.J Virol. 2008 Mar;82(6):2966-74. doi: 10.1128/JVI.02060-07. Epub 2007 Dec 26. J Virol. 2008. PMID: 18160430 Free PMC article.
-
A molecular understanding of alphavirus entry and antibody protection.Nat Rev Microbiol. 2023 Jun;21(6):396-407. doi: 10.1038/s41579-022-00825-7. Epub 2022 Dec 6. Nat Rev Microbiol. 2023. PMID: 36474012 Free PMC article. Review.
-
Fusion of mApple and Venus fluorescent proteins to the Sindbis virus E2 protein leads to different cell-binding properties.Virus Res. 2013 Nov 6;177(2):138-46. doi: 10.1016/j.virusres.2013.07.014. Epub 2013 Jul 31. Virus Res. 2013. PMID: 23916968 Free PMC article.
-
Winter ecology of Buggy Creek virus (Togaviridae, Alphavirus) in the Central Great Plains.Vector Borne Zoonotic Dis. 2010 May;10(4):355-63. doi: 10.1089/vbz.2009.0031. Vector Borne Zoonotic Dis. 2010. PMID: 19725760 Free PMC article.
-
A structural and functional perspective of alphavirus replication and assembly.Future Microbiol. 2009 Sep;4(7):837-56. doi: 10.2217/fmb.09.59. Future Microbiol. 2009. PMID: 19722838 Free PMC article. Review.
References
-
- Chikkanna-Gowda CP, Sheahan BJ, Fleeton MN, Atkins GJ. Regression of mouse tumours and inhibition of metastases following administration of a Semliki Forest virus vector with enhanced expression of IL-12. Gene Ther. 2005;12(16):1253–63. - PubMed
-
- Choi HK, Tong L, Minor W, Dumas P, Boege U, Rossmann MG, Wengler G. Structure of Sindbis virus core protein reveals a chymotrypsin-like serine proteinase and the organization of the virion. Nature. 1991;354:37–43. - PubMed
-
- Davis NL, Pence DF, Meyer WJ, Schmaljohn AL, Johnston RE. Alternative forms of a strain-specific neutralizing antigenic site on the Sindbis virus E2 glycoprotein. Virology. 1987;161(1):101–8. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

