Complete genome sequence of the prototype lactic acid bacterium Lactococcus lactis subsp. cremoris MG1363
- PMID: 17307855
- PMCID: PMC1855848
- DOI: 10.1128/JB.01768-06
Complete genome sequence of the prototype lactic acid bacterium Lactococcus lactis subsp. cremoris MG1363
Abstract
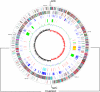
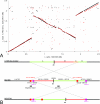
Lactococcus lactis is of great importance for the nutrition of hundreds of millions of people worldwide. This paper describes the genome sequence of Lactococcus lactis subsp. cremoris MG1363, the lactococcal strain most intensively studied throughout the world. The 2,529,478-bp genome contains 81 pseudogenes and encodes 2,436 proteins. Of the 530 unique proteins, 47 belong to the COG (clusters of orthologous groups) functional category "carbohydrate metabolism and transport," by far the largest category of novel proteins in comparison with L. lactis subsp. lactis IL1403. Nearly one-fifth of the 71 insertion elements are concentrated in a specific 56-kb region. This integration hot-spot region carries genes that are typically associated with lactococcal plasmids and a repeat sequence specifically found on plasmids and in the "lateral gene transfer hot spot" in the genome of Streptococcus thermophilus. Although the parent of L. lactis MG1363 was used to demonstrate lysogeny in Lactococcus, L. lactis MG1363 carries four remnant/satellite phages and two apparently complete prophages. The availability of the L. lactis MG1363 genome sequence will reinforce its status as the prototype among lactic acid bacteria through facilitation of further applied and fundamental research.
Figures

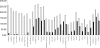
Similar articles
-
Physical and genetic map of the Lactococcus lactis subsp. cremoris MG1363 chromosome: comparison with that of Lactococcus lactis subsp. lactis IL 1403 reveals a large genome inversion.J Bacteriol. 1995 May;177(10):2840-50. doi: 10.1128/jb.177.10.2840-2850.1995. J Bacteriol. 1995. PMID: 7751295 Free PMC article.
-
Expression of prophage-encoded endolysins contributes to autolysis of Lactococcus lactis.Appl Microbiol Biotechnol. 2017 Feb;101(3):1099-1110. doi: 10.1007/s00253-016-7822-z. Epub 2016 Sep 22. Appl Microbiol Biotechnol. 2017. PMID: 27660179 Free PMC article.
-
Cloning, sequencing and comparison of three lactococcal L-lactate dehydrogenase genes.Microbiology (Reading). 1994 Jun;140 ( Pt 6):1301-5. doi: 10.1099/00221287-140-6-1301. Microbiology (Reading). 1994. PMID: 8081494
-
Low-redundancy sequencing of the entire Lactococcus lactis IL1403 genome.Antonie Van Leeuwenhoek. 1999 Jul-Nov;76(1-4):27-76. Antonie Van Leeuwenhoek. 1999. PMID: 10532372 Review.
-
Genomic organization of lactic acid bacteria.Antonie Van Leeuwenhoek. 1996 Oct;70(2-4):161-83. doi: 10.1007/BF00395932. Antonie Van Leeuwenhoek. 1996. PMID: 8879406 Review.
Cited by
-
KOMODO: a web tool for detecting and visualizing biased distribution of groups of homologous genes in monophyletic taxa.Nucleic Acids Res. 2012 Jul;40(Web Server issue):W491-7. doi: 10.1093/nar/gks490. Epub 2012 Jun 6. Nucleic Acids Res. 2012. PMID: 22675073 Free PMC article.
-
Inactivation of an iron transporter in Lactococcus lactis results in resistance to tellurite and oxidative stress.Appl Environ Microbiol. 2007 Oct;73(19):6144-9. doi: 10.1128/AEM.00413-07. Epub 2007 Aug 3. Appl Environ Microbiol. 2007. PMID: 17675432 Free PMC article.
-
Tracing mother-infant transmission of bacteriophages by means of a novel analytical tool for shotgun metagenomic datasets: METAnnotatorX.Microbiome. 2018 Aug 20;6(1):145. doi: 10.1186/s40168-018-0527-z. Microbiome. 2018. PMID: 30126456 Free PMC article.
-
Design and Expression of Specific Hybrid Lantibiotics Active Against Pathogenic Clostridium spp.Front Microbiol. 2019 Sep 24;10:2154. doi: 10.3389/fmicb.2019.02154. eCollection 2019. Front Microbiol. 2019. PMID: 31616392 Free PMC article.
-
Genome sequences of Lactococcus lactis MG1363 (revised) and NZ9000 and comparative physiological studies.J Bacteriol. 2010 Nov;192(21):5806-12. doi: 10.1128/JB.00533-10. Epub 2010 Jul 16. J Bacteriol. 2010. PMID: 20639323 Free PMC article.
References
-
- Altermann, E., W. M. Russell, M. A. Azcarate-Peril, R. Barrangou, B. L. Buck, O. McAuliffe, N. Souther, A. Dobson, T. Duong, M. Callanan, S. Lick, A. Hamrick, R. Cano, and T. R. Klaenhammer. 2005. Complete genome sequence of the probiotic lactic acid bacterium Lactobacillus acidophilus NCFM. Proc. Natl. Acad. Sci. USA 102:3906-3912. - PMC - PubMed
-
- Audouy, S. A. L., M. L. van Roosmalen, J. Neef, R. Kanninga, E. Post, M. van Deemter, H. Metselaar, S. van Selm, G. T. Robillard, K. J. Leenhouts, and P. W. M. Hermans. 2006. Lactococcus lactis GEM particles displaying pneumococcal antigens induce local and systemic immune responses following intranasal immunization. Vaccine 24:5434-5441. - PubMed
-
- Badger, J. H., and G. J. Olsen. 1999. CRITICA: coding region identification tool invoking comparative analysis. Mol. Biol. Evol. 16:512-524. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

