Chlamydial Hsp60-2 is iron responsive in Chlamydia trachomatis serovar E-infected human endometrial epithelial cells in vitro
- PMID: 17307941
- PMCID: PMC1865735
- DOI: 10.1128/IAI.01465-06
Chlamydial Hsp60-2 is iron responsive in Chlamydia trachomatis serovar E-infected human endometrial epithelial cells in vitro
Abstract
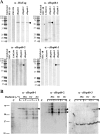
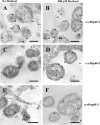
Chlamydial 60-kDa heat shock proteins (cHsp60s) are known to play a prominent role in the immunopathogenesis of disease. It is also known that several stress-inducing growth conditions, such as heat, iron deprivation, or exposure to gamma interferon, result in the development of persistent chlamydial forms that often exhibit enhanced expression of cHsp60. We have shown previously that the expression of cHsp60 is greatly enhanced in Chlamydia trachomatis serovar E propagated in an iron-deficient medium. The objective of this work was to determine which single cHsp60 or combination of the three cHsp60 homologs encoded by this organism responds to iron limitation. Using monospecific polyclonal peptide antisera that recognize only cHsp60-1, cHsp60-2, or cHsp60-3, we found that expression of cHsp60-2 is responsive to iron deprivation. Overall, our studies suggest that the expression of cHsp60 homologs differs among the mechanisms currently known to induce persistence.
Figures

References
-
- Andrews, N. C. 2000. Iron homeostasis: insights from genetics and animal models. Nat. Rev. 1:208-216. - PubMed
-
- Beatty, W. L., G. I. Byrne, and R. P. Morrison. 1994. Repeated and persistent infection with Chlamydia and the development of chronic inflammation and disease. Trends Microbiol. 2:94-98. - PubMed
-
- Belland, R. J., L. Rose, and Y. AbdelRahman. 2006. Promoter mapping of coding and non-coding RNAs using an intergenic microarray for Chlamydia trachomatis, p. 17-20. In M. Chernesky, H. Caldwell, G. Christiansen, I. N. Clarke, B. Kaltenboek, C. Knirsch, C.-C. Kuo, J. Mahony, R. G. Rank, P. Saikku, J. Schachter, W. E. Stamm, R. S. Stephens, J. T. Summersgill, P. Timms, and P. B. Wyrick (ed.), Proceedings of the 11th International Symposium on Human Chlamydial Infections, Niagara-on-the-Lake, Ontario, Canada. International Chlamydia Symposium, San Francisco, CA.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

