Common structural requirements for heptahelical domain function in class A and class C G protein-coupled receptors
- PMID: 17310064
- PMCID: PMC2565688
- DOI: 10.1074/jbc.M611071200
Common structural requirements for heptahelical domain function in class A and class C G protein-coupled receptors
Abstract
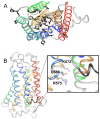
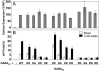
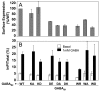
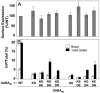
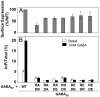
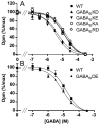
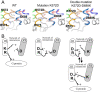
G protein-coupled receptors (GPCRs) are key players in cell communication. Several classes of such receptors have been identified. Although all GPCRs possess a heptahelical domain directly activating G proteins, important structural and sequence differences within receptors from different classes suggested distinct activation mechanisms. Here we show that highly conserved charged residues likely involved in an interaction network between transmembrane domains (TM) 3 and 6 at the cytoplasmic side of class C GPCRs are critical for activation of the gamma-aminobutyric acid type B receptor. Indeed, the loss of function resulting from the mutation of the conserved lysine residue into aspartate or glutamate in the TM3 of gamma-aminobutyric acid type B(2) can be partly rescued by mutating the conserved acidic residue of TM6 into either lysine or arginine. In addition, mutation of the conserved lysine into an acidic residue leads to a nonfunctional receptor that displays a high agonist affinity. This is reminiscent of a similar ionic network that constitutes a lock stabilizing the inactive state of many class A rhodopsin-like GPCRs. These data reveal that despite their original structure, class C GPCRs share with class A receptors at least some common structural feature controlling G protein activation.
Figures


References
-
- Bockaert J, Claeysen S, Becamel C, Pinloche S, Dumuis A. Int Rev Cytol. 2002;212:63–132. - PubMed
-
- Karnik SS, Gogonea C, Patil S, Saad Y, Takezako T. Trends Endocrinol Metab. 2003;14:431–437. - PubMed
-
- Gether U. Endocr Rev. 2000;21:90–113. - PubMed
-
- Palczewski K, Kumasaka T, Hori T, Behnke CA, Motoshima H, Fox BA, Le Trong I, Teller DC, Okada T, Stenkamp RE, Yamamoto M, Miyano M. Science. 2000;289:739–745. - PubMed
-
- Fredriksson R, Lagerstrom MC, Lundin LG, Schioth HB. Mol Pharmacol. 2003;63:1256–1272. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

