Understanding and using the meaning of statements in a bio-ontology: recasting the Gene Ontology in OWL
- PMID: 17311682
- PMCID: PMC1819394
- DOI: 10.1186/1471-2105-8-57
Understanding and using the meaning of statements in a bio-ontology: recasting the Gene Ontology in OWL
Abstract
The bio-ontology community falls into two camps: first we have biology domain experts, who actually hold the knowledge we wish to capture in ontologies; second, we have ontology specialists, who hold knowledge about techniques and best practice on ontology development. In the bio-ontology domain, these two camps have often come into conflict, especially where pragmatism comes into conflict with perceived best practice. One of these areas is the insistence of computer scientists on a well-defined semantic basis for the Knowledge Representation language being used. In this article, we will first describe why this community is so insistent. Second, we will illustrate this by examining the semantics of the Web Ontology Language and the semantics placed on the Directed Acyclic Graph as used by the Gene Ontology. Finally we will reconcile the two representations, including the broader Open Biomedical Ontologies format. The ability to exchange between the two representations means that we can capitalise on the features of both languages. Such utility can only arise by the understanding of the semantics of the languages being used. By this illustration of the usefulness of a clear, well-defined language semantics, we wish to promote a wider understanding of the computer science perspective amongst potential users within the biological community.
Figures
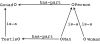

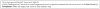
References
-
- OWL Web Ontology Language Guide http://www.w3.org/TR/owl-guide
-
- Horrocks I, Patel-Schneider PF, van Harmelen F. From SHIQ and RDF to OWL: The Making of a Web Ontology Language. Journal of Web Semantics. 2003;1:7–26.
-
- Open Biomedical Ontology http://obo.sourceforge.net
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

