Rapid, accurate, computational discovery of Rho-independent transcription terminators illuminates their relationship to DNA uptake
- PMID: 17313685
- PMCID: PMC1852404
- DOI: 10.1186/gb-2007-8-2-r22
Rapid, accurate, computational discovery of Rho-independent transcription terminators illuminates their relationship to DNA uptake
Abstract
Background: In many prokaryotes, transcription of DNA to RNA is terminated by a thymine-rich stretch of DNA following a hairpin loop. Detecting such Rho-independent transcription terminators can shed light on the organization of bacterial genomes and can improve genome annotation. Previous computational methods to predict Rho-independent terminators have been slow or limited in the organisms they consider.
Results: We describe TransTermHP, a new computational method to rapidly and accurately detect Rho-independent transcription terminators. We predict the locations of terminators in 343 prokaryotic genomes, representing the largest collection of predictions available. In Bacillus subtilis, we can detect 93% of known terminators with a false positive rate of just 6%, comparable to the best-known methods. Outside the Firmicutes division, we find that Rho-independent termination plays a large role in the Neisseria and Vibrio genera, the Pasteurellaceae (including the Haemophilus genus) and several other species. In Neisseria and Pasteurellaceae, terminator hairpins are frequently formed by closely spaced, complementary instances of exogenous DNA uptake signal sequences. We quantify the propensity for terminators to include these sequences. In the process, we provide the first discussion of potential uptake signals in Haemophilus ducreyi and Mannheimia succiniciproducens, and we discuss the preference for a particular configuration of uptake signal sequences within terminators.
Conclusion: Our new fast and accurate method for detecting transcription terminators has allowed us to identify and analyze terminators in many new genomes and to identify DNA uptake signal sequences in several species where they have not been previously reported. Our software and predictions are freely available.
Figures
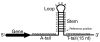


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

