Distinct energetics and closing pathways for DNA polymerase beta with 8-oxoG template and different incoming nucleotides
- PMID: 17313689
- PMCID: PMC1819382
- DOI: 10.1186/1472-6807-7-7
Distinct energetics and closing pathways for DNA polymerase beta with 8-oxoG template and different incoming nucleotides
Abstract
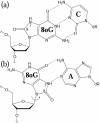
Background: 8-Oxoguanine (8-oxoG) is a common oxidative lesion frequently encountered by DNA polymerases such as the repair enzyme DNA polymerase beta (pol beta). To interpret in atomic and energetic detail how pol beta processes 8-oxoG, we apply transition path sampling to delineate closing pathways of pol beta 8-oxoG complexes with dCTP and dATP incoming nucleotides and compare the results to those of the nonlesioned G:dCTP and G:dATPanalogues.
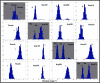
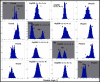
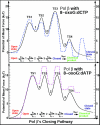
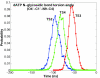
Results: Our analyses show that the closing pathways of the 8-oxoG complexes are different from one another and from the nonlesioned analogues in terms of the individual transition states along each pathway, associated energies, and the stability of each pathway's closed state relative to the corresponding open state. In particular, the closed-to-open state stability difference in each system establishes a hierarchy of stability (from high to low) as G:C > 8-oxoG:C > 8-oxoG:A > G:A, corresponding to -3, -2, 2, 9 kBT, respectively. This hierarchy of closed state stability parallels the experimentally observed processing efficiencies for the four pairs. Network models based on the calculated rate constants in each pathway indicate that the closed species are more populated than the open species for 8-oxoG:dCTP, whereas the opposite is true for 8-oxoG:dATP.
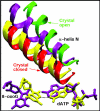
Conclusion: These results suggest that the lower insertion efficiency (larger Km) for dATP compared to dCTP opposite 8-oxoG is caused by a less stable closed-form of pol beta, destabilized by unfavorable interactions between Tyr271 and the mispair. This stability of the closed vs. open form can also explain the higher insertion efficiency for 8-oxoG:dATP compared to the nonlesioned G:dATP pair, which also has a higher overall conformational barrier. Our study offers atomic details of the complexes at different states, in addition to helping interpret the different insertion efficiencies of dATP and dCTP opposite 8-oxoG and G.
Figures

Similar articles
-
Differing conformational pathways before and after chemistry for insertion of dATP versus dCTP opposite 8-oxoG in DNA polymerase beta.Biophys J. 2007 May 1;92(9):3063-70. doi: 10.1529/biophysj.106.092106. Epub 2007 Feb 9. Biophys J. 2007. PMID: 17293403 Free PMC article.
-
How DNA polymerase X preferentially accommodates incoming dATP opposite 8-oxoguanine on the template.Biophys J. 2013 Dec 3;105(11):2559-68. doi: 10.1016/j.bpj.2013.10.014. Biophys J. 2013. PMID: 24314086 Free PMC article.
-
Analysis of nucleotide insertion and extension at 8-oxo-7,8-dihydroguanine by replicative T7 polymerase exo- and human immunodeficiency virus-1 reverse transcriptase using steady-state and pre-steady-state kinetics.Biochemistry. 1997 May 27;36(21):6475-87. doi: 10.1021/bi9627267. Biochemistry. 1997. PMID: 9174365
-
DNA polymerase structure-based insight on the mutagenic properties of 8-oxoguanine.Mutat Res. 2010 Nov 28;703(1):18-23. doi: 10.1016/j.mrgentox.2010.07.013. Epub 2010 Aug 7. Mutat Res. 2010. PMID: 20696268 Free PMC article. Review.
-
A Role for N6-Methyladenine in DNA Damage Repair.Trends Biochem Sci. 2021 Mar;46(3):175-183. doi: 10.1016/j.tibs.2020.09.007. Epub 2020 Oct 16. Trends Biochem Sci. 2021. PMID: 33077363 Free PMC article. Review.
Cited by
-
Differing conformational pathways before and after chemistry for insertion of dATP versus dCTP opposite 8-oxoG in DNA polymerase beta.Biophys J. 2007 May 1;92(9):3063-70. doi: 10.1529/biophysj.106.092106. Epub 2007 Feb 9. Biophys J. 2007. PMID: 17293403 Free PMC article.
-
DNA polymerase family X: function, structure, and cellular roles.Biochim Biophys Acta. 2010 May;1804(5):1136-50. doi: 10.1016/j.bbapap.2009.07.008. Epub 2009 Jul 23. Biochim Biophys Acta. 2010. PMID: 19631767 Free PMC article. Review.
-
Role of Electron-Driven Proton-Transfer Processes in the Ultrafast Deactivation of Photoexcited Anionic 8-oxoGuanine-Adenine and 8-oxoGuanine-Cytosine Base Pairs.Molecules. 2017 Jan 14;22(1):135. doi: 10.3390/molecules22010135. Molecules. 2017. PMID: 28098833 Free PMC article.
-
Mismatched base-pair simulations for ASFV Pol X/DNA complexes help interpret frequent G*G misincorporation.J Mol Biol. 2008 Dec 31;384(5):1086-97. doi: 10.1016/j.jmb.2008.10.025. Epub 2008 Oct 17. J Mol Biol. 2008. PMID: 18955064 Free PMC article.
-
"Gate-keeper" residues and active-site rearrangements in DNA polymerase μ help discriminate non-cognate nucleotides.PLoS Comput Biol. 2013;9(5):e1003074. doi: 10.1371/journal.pcbi.1003074. Epub 2013 May 23. PLoS Comput Biol. 2013. PMID: 23717197 Free PMC article.
References
-
- Mol CD, Parikh SS, Putnam CD, Lo TP, Tainer JA. DNA repair mechanisms for the recognition and removal of damaged DNA bases. Annu Rev Biophys Biomol Struct. 1999;28:101–128. - PubMed
-
- Lindhal T. Instability and decay of the primary structure of DNA. Nature. 1993;362:709–715. - PubMed
-
- Ames BN. Dietary carcinogens and anticarcinogens - oxygen radicals and degenerative diseases. Science. 1983;221:1256–1264. - PubMed
-
- Finkel T, Holbrook NJ. Oxidants, oxidative stress and the biology of ageing. Nature. 2000;408:239–247. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

