Development of a real-time PCR probe for quantification of the heterotrophic dinoflagellate Cryptoperidiniopsis brodyi (Dinophyceae) in environmental samples
- PMID: 17322326
- PMCID: PMC1855596
- DOI: 10.1128/AEM.02389-06
Development of a real-time PCR probe for quantification of the heterotrophic dinoflagellate Cryptoperidiniopsis brodyi (Dinophyceae) in environmental samples
Abstract
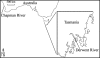
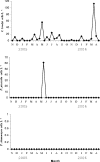
A TaqMan format real-time PCR probe was developed against the internal transcribed spacer 2 ribosomal DNA region for the specific detection and quantification of Cryptoperidiniopsis brodyi in environmental samples. The assay specificity was confirmed by testing against related dinoflagellates and verified by sequencing PCR amplicons from natural water samples. Phylogenetic analysis of the sequenced environmental samples also showed that this assay is specific to C. brodyi. The C. brodyi-specific assay was used in conjunction with Pfiesteria piscicida- and Pfiesteria shumwayae-specific real-time PCR assays to investigate the temporal variations of C. brodyi, P. piscicida, and P. shumwayae abundance in the Derwent estuary, Tasmania. The 18-month field survey from November 2004 to April 2006 revealed that C. brodyi occurred in all seasons at very low densities, mostly below 25 cells liter(-1), with higher abundance (maximum, 112 cells liter(-1)) in April and May. P. piscicida was detected only once, in May 2005 at 60 cells liter(-1). P. shumwayae was not detected during the survey.
Figures


References
-
- Bolch, C. J. S. 2001. PCR protocol for genetic identification of dinoflagellates directly from single cysts and plankton cells. Phycologia 40:162-168.
-
- Bowers, H. A., T. M. Trice, R. E. Magien, D. M. Goshorn, B. Michael, E. F. Schaefer, P. A. Rublee, and D. W. Oldach. 2006. Detection of Pfiesteria spp. by PCR in surface sediments collected from Chesapeake Bay tributaries (Maryland). Harmful Algae 5:342-351.
-
- Burkholder, J. M., and H. B. Glasgow. 1995. Interactions of a toxic estuarine dinoflagellate with microbial predations and prey. Arch. Protistenkd. 145:177-188.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Miscellaneous

