(Un)confined diffusion of CD59 in the plasma membrane determined by high-resolution single molecule microscopy
- PMID: 17325009
- PMCID: PMC1853144
- DOI: 10.1529/biophysj.106.095398
(Un)confined diffusion of CD59 in the plasma membrane determined by high-resolution single molecule microscopy
Abstract
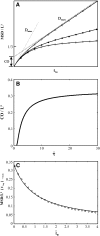
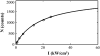
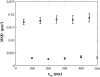
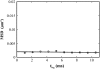
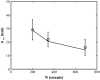
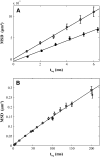
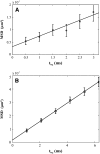
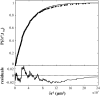
There has been emerging interest whether plasma membrane constituents are moving according to free Brownian motion or hop diffusion. In the latter model, lipids, lipid-anchored proteins, and transmembrane proteins would be transiently confined to periodic corrals in the cell membrane, which are structured by the underlying membrane skeleton. Because this model is based exclusively on results provided by one experimental strategy--high-resolution single particle tracking--we attempted in this study to confirm or amend it using a complementary technique. We developed a novel strategy that employs single molecule fluorescence microscopy to detect confinements to free diffusion of CD59--a GPI-anchored protein--in the plasma membrane of living T24 (ECV) cells. With this method, minimum invasive labeling via fluorescent Fab fragments was sufficient to measure the lateral motion of individual protein molecules on a millisecond timescale, yielding a positional accuracy down to 22 nm. Although no hop diffusion was directly observable, based on a full analytical description our results provide upper boundaries for confinement size and strength.
Figures


References
-
- Sharma, P., R. Varma, R.C. Sarasij, Ira, K. Gousset, G. Krishnamoorthy, M. Rao, and S. Mayor. 2004. Nanoscale organization of multiple GPI-anchored proteins in living cell membranes. Cell 116:577–589. - PubMed
-
- Simons, K., and E. Ikonen. 1997. Functional rafts in cell membranes. Nature. 387:569–572. - PubMed
-
- Munro, S. 2003. Lipid rafts: elusive or illusive? Cell. 115:377–388. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

