Tissue and subcellular distribution of CLIC1
- PMID: 17326840
- PMCID: PMC1820597
- DOI: 10.1186/1471-2121-8-8
Tissue and subcellular distribution of CLIC1
Abstract
Background: CLIC1 is a chloride channel whose cellular role remains uncertain. The distribution of CLIC1 in normal tissues is largely unknown and conflicting data have been reported regarding the cellular membrane fraction in which CLIC1 resides.
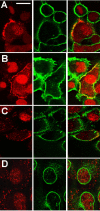
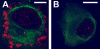
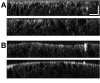
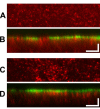
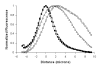
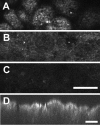
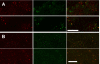
Results: New antisera to CLIC1 were generated and were found to be sensitive and specific for detecting this protein. These antisera were used to investigate the distribution of CLIC1 in mouse tissue sections and three cultured cell lines. We find CLIC1 is expressed in the apical domains of several simple columnar epithelia including glandular stomach, small intestine, colon, bile ducts, pancreatic ducts, airway, and the tail of the epididymis, in addition to the previously reported renal proximal tubule. CLIC1 is expressed in a non-polarized distribution in the basal epithelial cell layer of the stratified squamous epithelium of the upper gastrointesitinal tract and the basal cells of the epididymis, and is present diffusely in skeletal muscle. Distribution of CLIC1 was examined in Panc1 cells, a relatively undifferentiated, non-polarized human cell line derived from pancreatic cancer, and T84 cells, a human colon cancer cell line which can form a polarized epithelium that is capable of regulated chloride transport. Digitonin extraction was used to distinguish membrane-inserted CLIC1 from the soluble cytoplasmic form of the protein. We find that digitonin-resistant CLIC1 is primarily present in the plasma membrane of Panc1 cells. In T84 cells, we find digitonin-resistant CLIC1 is present in an intracellular compartment which is concentrated immediately below the apical plasma membrane and the extent of apical polarization is enhanced with forskolin, which activates transepithelial chloride transport and apical membrane traffic in these cells. The sub-apical CLIC1 compartment was further characterized in a well-differentiated mouse renal proximal tubule cell line. The distribution of CLIC1 was found to overlap that of megalin and the sodium-phosphate cotransporter, NaPi-II, which are markers of the apical endocytic/recycling compartment in proximal tubule.
Conclusion: The cell and tissue specific patterns of CLIC1 expression suggest it may play distinct roles in different cell types. In certain polarized columnar epithelia, it may play a role in apical membrane recycling.
Figures




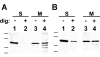

References
-
- Harrop SJ, DeMaere MZ, Fairlie WD, Reztsova T, Valenzuela SM, Mazzanti M, Tonini R, Qiu MR, Jankova L, Warton K, Bauskin AR, Wu WM, Pankhurst S, Campbell T, Breit SN, Cumi PM. Crystal structure of a soluble form of the intracellular chloride ion channel CLIC1 (NCC27) at 1.4-A resolution. J Biol Chem. 2001;276:44993–44500. doi: 10.1074/jbc.M107804200. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

