IrAE: an asparaginyl endopeptidase (legumain) in the gut of the hard tick Ixodes ricinus
- PMID: 17336985
- PMCID: PMC2587490
- DOI: 10.1016/j.ijpara.2006.12.020
IrAE: an asparaginyl endopeptidase (legumain) in the gut of the hard tick Ixodes ricinus
Abstract
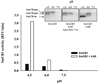
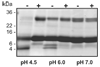
Ticks are ectoparasitic blood-feeders and important vectors for pathogens including arboviruses, rickettsiae, spirochetes and protozoa. As obligate blood-feeders, one possible strategy to retard disease transmission is disruption of the parasite's ability to digest host proteins. However, the constituent peptidases in the parasite gut and their potential interplay in the digestion of the blood meal are poorly understood. We have characterised a novel asparaginyl endopeptidase (legumain) from the hard tick Ixodes ricinus (termed IrAE), which we believe is the first such characterisation of a clan CD family C13 cysteine peptidase (protease) in arthropods. By RT-PCR of different tissues, IrAE mRNA was only expressed in the tick gut. Indirect immunofluorescence and EM localised IrAE in the digestive vesicles of gut cells and within the peritrophic matrix. IrAE was functionally expressed in Pichia pastoris and reacted with a specific peptidyl fluorogenic substrate, and acyloxymethyl ketone and aza-asparagine Michael acceptor inhibitors. IrAE activity was unstable at pH > or = 6.0 and was shown to have a strict specificity for asparagine at P1 using a positional scanning synthetic combinatorial library. The enzyme hydrolyzed protein substrates with a pH optimum of 4.5, consistent with the pH of gut cell digestive vesicles. Thus, IrAE cleaved the major protein of the blood meal, hemoglobin, to a predominant peptide of 4kDa. Also, IrAE trans-processed and activated the zymogen form of Schistosoma mansoni cathepsin B1 -- an enzyme contributing to hemoglobin digestion in the gut of that bloodfluke. The possible functions of IrAE in the gut digestive processes of I. ricinus are compared with those suggested for other hematophagous parasites.
Figures







References
-
- Agyei AD, Runham NW, Blackstock N. Histochemical changes in the midgut of two ixodid tick species Boophilus microplus and Rhipicephalus appendiculatus during digestion of the blood meal. Exp. Appl. Acarol. 1992;13:187–212. - PubMed
-
- Bendtsen JD, Nielsen H, von Heijne G, Brunak S. Improved prediction of signal peptides: SignalP 3.0. J. Mol. Biol. 2004;340:783–795. - PubMed
-
- Boldbaatar D, Sikalizyo Sikasunge C, Battsetseg B, Xuan X, Fujisaki K. Molecular cloning and functional characterization of an aspartic protease from the hard tick Haemaphysalis longicornis. Insect Biochem. Mol. Biol. 2006;36:25–36. - PubMed
-
- Caffrey CR, Mathieu MA, Gaffney AM, Salter JP, Sajid M, Lucas KD, Franklin C, Bogyo M, McKerrow JH. Identification of a cDNA encoding an active asparaginyl endopeptidase of Schistosoma mansoni and its expression in Pichia pastoris. FEBS Lett. 2000;466:244–248. - PubMed
-
- Caffrey CR, McKerrow JH, Salter JP, Sajid M. Blood ‘n’ guts: an update on schistosome digestive peptidases. Trends Parasitol. 2004;20:241–248. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

