A unified view of base excision repair: lesion-dependent protein complexes regulated by post-translational modification
- PMID: 17337257
- PMCID: PMC1995033
- DOI: 10.1016/j.dnarep.2007.01.009
A unified view of base excision repair: lesion-dependent protein complexes regulated by post-translational modification
Abstract
Base excision repair (BER) proteins act upon a significantly broad spectrum of DNA lesions that result from endogenous and exogenous sources. Multiple sub-pathways of BER (short-path or long-patch) and newly designated DNA repair pathways (e.g., SSBR and NIR) that utilize BER proteins complicate any comprehensive understanding of BER and its role in genome maintenance, chemotherapeutic response, neuro-degeneration, cancer or aging. Herein, we propose a unified model of BER, comprised of three functional processes: Lesion Recognition/Strand Scission, Gap Tailoring and DNA Synthesis/Ligation, each represented by one or more multi-protein complexes and coordinated via the XRCC1/DNA Ligase III and PARP1 scaffold proteins. BER therefore may be represented by a series of repair complexes that assemble at the site of the DNA lesion and mediates repair in a coordinated fashion involving protein-protein interactions that dictate subsequent steps or sub-pathway choice. Complex formation is influenced by post-translational protein modifications that arise from the cellular state or the DNA damage response, providing an increase in specificity and efficiency to the BER pathway. In this review, we have summarized the reported BER protein-protein interactions and protein post-translational modifications and discuss the impact on DNA repair capacity and complex formation.
Figures
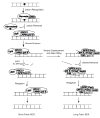

References
-
- Hitomi K, Iwai S, Tainer JA. The intricate structural chemistry of base excision repair machinery: Implications for DNA damage recognition, removal, and repair. DNA Repair (Amst) 2007 In Press. - PubMed
-
- Wilson SH, Kunkel TA. Passing the baton in base excision repair. Nature Structural Biology. 2000;7:176–178. - PubMed
-
- Nakamura J, Walker VE, Upton PB, Chiang SY, Kow YW, Swenberg JA. Highly sensitive apurinic/apyrimidinic site assay can detect spontaneous and chemically induced depurination under physiological conditions. Cancer Research. 1998;58:222–225. - PubMed
-
- Lindahl T, Wood RD. Quality control by DNA repair. Science. 1999;286:1897–1905. - PubMed
-
- Lindahl T. Instability and decay of the primary structure of DNA. Nature. 1993;362:709–715. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

