Knowledge-based localization of hippocampus in human brain MRI
- PMID: 17339035
- PMCID: PMC4502929
- DOI: 10.1016/j.compbiomed.2006.12.010
Knowledge-based localization of hippocampus in human brain MRI
Abstract
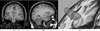
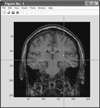
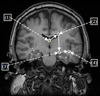
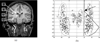
We present a novel and efficient method for localization of human brain structures such as hippocampus. Landmark localization is important for segmentation and registration. This method follows a statistical roadmap, consisting of anatomical landmarks, to reach the desired structures. Using a set of desired and undesired landmarks, identified on a training set, we estimate Gaussian models and determine optimal search areas for desired landmarks. The statistical models form a set of rules to evaluate the extracted landmarks during the search procedure. When applied on 900 MR images of 10 epileptic patients, this method demonstrated an overall success rate of 83%.
Figures















Similar articles
-
A model-based, semi-global segmentation approach for automatic 3-D point landmark localization in neuroimages.IEEE Trans Med Imaging. 2008 Aug;27(8):1034-44. doi: 10.1109/TMI.2008.915684. IEEE Trans Med Imaging. 2008. PMID: 18672421
-
Image registration of ex-vivo MRI to sparsely sectioned histology of hippocampal and neocortical temporal lobe specimens.Neuroimage. 2013 Dec;83:770-81. doi: 10.1016/j.neuroimage.2013.07.053. Epub 2013 Jul 26. Neuroimage. 2013. PMID: 23891884
-
Surface-based multi-template automated hippocampal segmentation: application to temporal lobe epilepsy.Med Image Anal. 2012 Oct;16(7):1445-55. doi: 10.1016/j.media.2012.04.008. Epub 2012 May 3. Med Image Anal. 2012. PMID: 22613821
-
Statistical validation of automatic methods for hippocampus segmentation in MR images of epileptic patients.Annu Int Conf IEEE Eng Med Biol Soc. 2014;2014:4707-10. doi: 10.1109/EMBC.2014.6944675. Annu Int Conf IEEE Eng Med Biol Soc. 2014. PMID: 25571043 Free PMC article.
-
Robust automated constellation-based landmark detection in human brain imaging.Neuroimage. 2018 Apr 15;170:471-481. doi: 10.1016/j.neuroimage.2017.04.012. Epub 2017 Apr 6. Neuroimage. 2018. PMID: 28392490 Free PMC article. Review.
Cited by
-
Hippocampus substructure segmentation using morphological vision transformer learning.Phys Med Biol. 2023 Dec 1;68(23):235013. doi: 10.1088/1361-6560/ad0d45. Phys Med Biol. 2023. PMID: 37972414 Free PMC article.
-
Superficial cortical landmarks for localization of the hippocampus: Application for temporal lobectomy and amygdalohippocampectomy.Surg Neurol Int. 2015 Feb 3;6:16. doi: 10.4103/2152-7806.150663. eCollection 2015. Surg Neurol Int. 2015. PMID: 25709853 Free PMC article.
-
External cortical landmarks for localization of the hippocampus: Application for temporal lobectomy and amygdalohippocampectomy.Surg Neurol Int. 2018 Aug 22;9:171. doi: 10.4103/sni.sni_446_17. eCollection 2018. Surg Neurol Int. 2018. PMID: 30210904 Free PMC article.
References
-
- Jack CR, Petersen RC, O’Brien PC, Tangalos EG. MR-based hippocampal volumetry in the diagnosis of Alzheimer’s disease. Neurology. 1992;42(1):183–188. - PubMed
-
- Cendes F, Andermann F, Gloor P, Evans A, Jones-Gotman M, Watson C, Melanson D, Olivier A, Peters T, Lopes-Cendes I, et al. MRI volumetric measurement of amygdala and hippocampus in temporal lobe epilepsy. Neurology. 1993;43(4):719–725. - PubMed
-
- Du AT, Schuff N, Amend D, Laakso MP, Hsu YY, Jagust WJ, Yaffe K, Kramer JH, Reed B, Norman D, Chui HC, Weiner MW. Magnetic resonance imaging of the entorhinal cortex and hippocampus in mild cognitive impairment and Alzheimer’s disease. J Neurol Neurosurg Psychiatry. 2001 Oct;71(4):441–447. - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

