AMP-forming acetyl-CoA synthetases in Archaea show unexpected diversity in substrate utilization
- PMID: 17350930
- PMCID: PMC2686389
- DOI: 10.1155/2006/738517
AMP-forming acetyl-CoA synthetases in Archaea show unexpected diversity in substrate utilization
Abstract
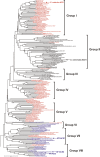
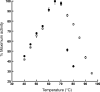
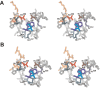
Adenosine monophosphate (AMP)-forming acetyl-CoA synthetase (ACS; acetate:CoA ligase (AMP-forming), EC 6.2.1.1) is a key enzyme for conversion of acetate to acetyl-CoA, an essential intermediate at the junction of anabolic and catabolic pathways. Phylogenetic analysis of putative short and medium chain acyl-CoA synthetase sequences indicates that the ACSs form a distinct clade from other acyl-CoA synthetases. Within this clade, the archaeal ACSs are not monophyletic and fall into three groups composed of both bacterial and archaeal sequences. Kinetic analysis of two archaeal enzymes, an ACS from Methanothermobacter thermautotrophicus (designated as MT-ACS1) and an ACS from Archaeoglobus fulgidus (designated as AF-ACS2), revealed that these enzymes have very different properties. MT-ACS1 has nearly 11-fold higher affinity and 14-fold higher catalytic efficiency with acetate than with propionate, a property shared by most ACSs. However, AF-ACS2 has only 2.3-fold higher affinity and catalytic efficiency with acetate than with propionate. This enzyme has an affinity for propionate that is almost identical to that of MT-ACS1 for acetate and nearly tenfold higher than the affinity of MT-ACS1 for propionate. Furthermore, MT-ACS1 is limited to acetate and propionate as acyl substrates, whereas AF-ACS2 can also utilize longer straight and branched chain acyl substrates. Phylogenetic analysis, sequence alignment and structural modeling suggest a molecular basis for the altered substrate preference and expanded substrate range of AF-ACS2 versus MT-ACS1.
Figures


Similar articles
-
Characterization of the acyl substrate binding pocket of acetyl-CoA synthetase.Biochemistry. 2006 Sep 26;45(38):11482-90. doi: 10.1021/bi061023e. Biochemistry. 2006. PMID: 16981708
-
AMP-forming acetyl-CoA synthetase from the extremely halophilic archaeon Haloarcula marismortui: purification, identification and expression of the encoding gene, and phylogenetic affiliation.Extremophiles. 2005 Oct;9(5):355-65. doi: 10.1007/s00792-005-0449-0. Epub 2005 Jun 10. Extremophiles. 2005. PMID: 15947865
-
Application of Acetyl-CoA synthetase from Methanothermobacter thermautotrophicus to non-native substrates.Enzyme Microb Technol. 2019 Sep;128:67-71. doi: 10.1016/j.enzmictec.2019.05.005. Epub 2019 May 14. Enzyme Microb Technol. 2019. PMID: 31186112
-
Acetyl-coenzyme A synthetase (AMP forming).Cell Mol Life Sci. 2004 Aug;61(16):2020-30. doi: 10.1007/s00018-004-3448-x. Cell Mol Life Sci. 2004. PMID: 15316652 Free PMC article. Review.
-
A link between transcription and intermediary metabolism: a role for Sir2 in the control of acetyl-coenzyme A synthetase.Curr Opin Microbiol. 2004 Apr;7(2):115-9. doi: 10.1016/j.mib.2004.02.005. Curr Opin Microbiol. 2004. PMID: 15063846 Review.
Cited by
-
Acetate Metabolism in Anaerobes from the Domain Archaea.Life (Basel). 2015 Jun 9;5(2):1454-71. doi: 10.3390/life5021454. Life (Basel). 2015. PMID: 26068860 Free PMC article. Review.
-
Genome-wide transcriptional response to silver stress in extremely halophilic archaeon Haloferax alexandrinus DSM 27206 T.BMC Microbiol. 2023 Dec 4;23(1):381. doi: 10.1186/s12866-023-03133-z. BMC Microbiol. 2023. PMID: 38049746 Free PMC article.
-
The Roles of Coenzyme A Binding Pocket Residues in Short and Medium Chain Acyl-CoA Synthetases.Life (Basel). 2023 Jul 28;13(8):1643. doi: 10.3390/life13081643. Life (Basel). 2023. PMID: 37629500 Free PMC article.
-
Protein adaptations in archaeal extremophiles.Archaea. 2013;2013:373275. doi: 10.1155/2013/373275. Epub 2013 Sep 16. Archaea. 2013. PMID: 24151449 Free PMC article. Review.
-
Physiological differentiation within a single-species biofilm fueled by serpentinization.mBio. 2011 Sep 1;2(4):e00127-11. doi: 10.1128/mBio.00127-11. Print 2011. mBio. 2011. PMID: 21791580 Free PMC article.
References
-
- Allen G.W.J., Zinder S.H. Methanogenesis from acetate by cell-free extracts of the thermophilic acetotrophic methanogen Methanothrixthermophila CALS-1. Arch. Microbiol. 1996;166:275–281. - PubMed
-
- Altschul S.F., Gish W., Miller W., Myers E.W., Lipman D.J. Basic local alignment search tool. J. Mol. Biol. 1990;215:403–410. - PubMed
-
- Anke H., Spector L.B. Evidence for an acetyl-enzyme intermediate in the action of acetyl-CoA synthetase. Biochem. Biophys. Res. Commun. 1975;67:767–773. - PubMed
-
- Armon A., Graur D., Ben-tal N. ConSurf: an algorithmic tool for the identification of functional regions in proteins by surface mapping of phylogenetic information. J. Mol. Biol. 2001;307:447–463. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

