Importance of calcium-binding site 2 in simian virus 40 infection
- PMID: 17360742
- PMCID: PMC1900253
- DOI: 10.1128/JVI.02195-06
Importance of calcium-binding site 2 in simian virus 40 infection
Abstract
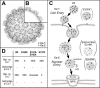
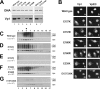
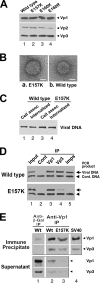
The exposure of molecular signals for simian virus 40 (SV40) cell entry and nuclear entry has been postulated to involve calcium coordination at two sites on the capsid made of Vp1. The role of calcium-binding site 2 in SV40 infection was examined by analyzing four single mutants of site 2, the Glu160Lys, Glu160Arg, Glu157Lys (E157K), and Glu157Arg mutants, and an E157K-E330K combination mutant. The last three mutants were nonviable. All mutants replicated viral DNA normally, and all except the last two produced particles containing all three capsid proteins and viral DNA. The defect of the site 1-site 2 E157K-E330K double mutant implies that at least one of the sites is required for particle assembly in vivo. The nonviable E157K particles, about 10% larger in diameter than the wild type, were able to enter cells but did not lead to T-antigen expression. Cell-internalized E157K DNA effectively coimmunoprecipitated with anti-Vp1 antibody, but little of the DNA did so with anti-Vp3 antibody, and none was detected in anti-importin immunoprecipitate. Yet, a substantial amount of Vp3 was present in anti-Vp1 immune complexes, suggesting that internalized E157K particles are ineffective at exposing Vp3. Our data show that E157K mutant infection is blocked at a stage prior to the interaction of the Vp3 nuclear localization signal with importins, consistent with a role for calcium-binding site 2 in postentry steps leading to the nuclear import of the infecting SV40.
Figures
Similar articles
-
Identification of amino acid residues within simian virus 40 capsid proteins Vp1, Vp2, and Vp3 that are required for their interaction and for viral infection.J Virol. 2006 Sep;80(18):8891-8. doi: 10.1128/JVI.00781-06. J Virol. 2006. PMID: 16940501 Free PMC article.
-
The simian virus 40 minor structural protein Vp3, but not Vp2, is essential for infectious virion formation.J Gen Virol. 2003 Aug;84(Pt 8):2111-2116. doi: 10.1099/vir.0.19287-0. J Gen Virol. 2003. PMID: 12867642
-
Simian virus 40 Vp1 DNA-binding domain is functionally separable from the overlapping nuclear localization signal and is required for effective virion formation and full viability.J Virol. 2001 Aug;75(16):7321-9. doi: 10.1128/JVI.75.16.7321-7329.2001. J Virol. 2001. PMID: 11462004 Free PMC article.
-
Host range analysis of simian virus 40, BK virus and chimaeric SV40/BKV: relative expression of large T-antigen and Vp1 in infected and transformed cells.Dev Biol Stand. 1998;94:191-205. Dev Biol Stand. 1998. PMID: 9776240 Review.
-
Simian virus 40 chromatin interaction with the capsid proteins.J Biomol Struct Dyn. 1983 Dec;1(3):689-704. doi: 10.1080/07391102.1983.10507475. J Biomol Struct Dyn. 1983. PMID: 6101085 Review.
Cited by
-
14-3-3zeta escorts CCTalpha for calcium-activated nuclear import in lung epithelia.FASEB J. 2010 Apr;24(4):1271-83. doi: 10.1096/fj.09-136044. Epub 2009 Dec 9. FASEB J. 2010. PMID: 20007511 Free PMC article.
-
Calcium bridge triggers capsid disassembly in the cell entry process of simian virus 40.J Biol Chem. 2009 Dec 11;284(50):34703-12. doi: 10.1074/jbc.M109.015107. Epub 2009 Oct 12. J Biol Chem. 2009. PMID: 19822519 Free PMC article.
-
Viral calciomics: interplays between Ca2+ and virus.Cell Calcium. 2009 Jul;46(1):1-17. doi: 10.1016/j.ceca.2009.05.005. Epub 2009 Jun 16. Cell Calcium. 2009. PMID: 19535138 Free PMC article. Review.
-
ERdj5 Reductase Cooperates with Protein Disulfide Isomerase To Promote Simian Virus 40 Endoplasmic Reticulum Membrane Translocation.J Virol. 2015 Sep;89(17):8897-908. doi: 10.1128/JVI.00941-15. Epub 2015 Jun 17. J Virol. 2015. PMID: 26085143 Free PMC article.
-
SV40 late protein VP4 forms toroidal pores to disrupt membranes for viral release.Biochemistry. 2013 Jun 4;52(22):3939-48. doi: 10.1021/bi400036z. Epub 2013 May 20. Biochemistry. 2013. PMID: 23651212 Free PMC article.
References
-
- Chang, D., C. Y. Fung, W. C. Ou, P. C. Chao, S. Y. Li, M. Wang, Y. L. Huang, T. Y. Tzeng, and R. T. Tsai. 1997. Self-assembly of the JC virus major capsid protein, VP1, expressed in insect cells. J. Gen. Virol. 78:1435-1439. - PubMed
-
- Chen, P. L., M. Wang, W. C. Ou, C. K. Lii, L. S. Chen, and D. Chang. 2001. Disulfide bonds stabilize JC virus capsid-like structure by protecting calcium ions from chelation. FEBS Lett. 500:109-113. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

