Functional analysis of cis- and trans-acting replication factors of porcine circovirus type 1
- PMID: 17360750
- PMCID: PMC1900318
- DOI: 10.1128/JVI.02420-06
Functional analysis of cis- and trans-acting replication factors of porcine circovirus type 1
Abstract
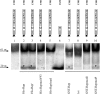
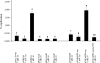
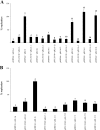
The replication proteins Rep and Rep' of porcine circovirus type 1 (PCV1) are both capable of introducing and resealing strand discontinuities at the viral origin of DNA replication in vitro underlying genome amplification by rolling-circle replication. The PCV1 origin of replication encompasses the minimal binding site (MBS) of the Rep and Rep' proteins and an inverted repeat with the potential to form a stem-loop. In this study, both elements of the PCV1 origin were demonstrated to be essential for viral replication in transfected cells. Furthermore, investigation of conserved amino acid motifs within Rep and Rep' proteins revealed that the mutation of motifs I, II, and III and of the GKS box interfered with viral replication. In vitro studies demonstrated that motifs I to III were essential for origin cleavage, while the GKS box was dispensable for the initiation of viral replication. A covalent link between Rep/Rep' and the DNA after origin cleavage was demonstrated, providing a mechanism for energy conservation for the termination of replication.
Figures

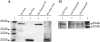


Similar articles
-
Demonstration of nicking/joining activity at the origin of DNA replication associated with the rep and rep' proteins of porcine circovirus type 1.J Virol. 2006 Jul;80(13):6225-34. doi: 10.1128/JVI.02506-05. J Virol. 2006. PMID: 16775310 Free PMC article.
-
Replication of porcine circovirus type 1 requires two proteins encoded by the viral rep gene.Virology. 2001 Jan 20;279(2):429-38. doi: 10.1006/viro.2000.0730. Virology. 2001. PMID: 11162799
-
Rep and Rep' protein of porcine circovirus type 1 bind to the origin of replication in vitro.Virology. 2001 Dec 5;291(1):152-60. doi: 10.1006/viro.2001.1203. Virology. 2001. PMID: 11878884
-
Porcine circovirus: transcription and DNA replication.Virus Res. 2012 Mar;164(1-2):46-53. doi: 10.1016/j.virusres.2011.10.012. Epub 2011 Oct 20. Virus Res. 2012. PMID: 22036834 Review.
-
Molecular biology of Porcine circovirus: analyses of gene expression and viral replication.Vet Microbiol. 2004 Feb 4;98(2):81-8. doi: 10.1016/j.vetmic.2003.10.014. Vet Microbiol. 2004. PMID: 14741119 Review.
Cited by
-
Replication of porcine circoviruses.Virol J. 2009 May 18;6:60. doi: 10.1186/1743-422X-6-60. Virol J. 2009. PMID: 19450240 Free PMC article. Review.
-
Diversity of CRESS DNA Viruses in Squamates Recapitulates Hosts Dietary and Environmental Sources of Exposure.Microbiol Spectr. 2022 Jun 29;10(3):e0078022. doi: 10.1128/spectrum.00780-22. Epub 2022 May 26. Microbiol Spectr. 2022. PMID: 35616383 Free PMC article.
-
Rapidly expanding genetic diversity and host range of the Circoviridae viral family and other Rep encoding small circular ssDNA genomes.Virus Res. 2012 Mar;164(1-2):114-21. doi: 10.1016/j.virusres.2011.11.021. Epub 2011 Dec 6. Virus Res. 2012. PMID: 22155583 Free PMC article. Review.
-
The fecal viral flora of wild rodents.PLoS Pathog. 2011 Sep;7(9):e1002218. doi: 10.1371/journal.ppat.1002218. Epub 2011 Sep 1. PLoS Pathog. 2011. PMID: 21909269 Free PMC article.
-
Multi-platform analysis reveals a complex transcriptome architecture of a circovirus.Virus Res. 2017 Jun 2;237:37-46. doi: 10.1016/j.virusres.2017.05.010. Epub 2017 May 24. Virus Res. 2017. PMID: 28549855 Free PMC article.
References
-
- Bassami, M. R., D. Berryman, G. E. Wilcox, and S. R. Raidal. 1998. Psittacine beak and feather disease virus nucleotide sequence analysis and its relationship to porcine circovirus, plant circoviruses, and chicken anaemia virus. Virology 249:453-459. - PubMed
-
- Cheung, A. K. 2005. Mutational analysis of the direct tandem repeat sequences at the origin of DNA replication of porcine circovirus type 1. Virology 339:192-199. - PubMed
-
- Danen-Van Oorschot, A. A., D. F. Fischer, J. M. Grimbergen, B. Klein, S. Zhuang, J. H. Falkenburg, C. Backendorf, P. H. Quax, A. J. Van der Eb, and M. H. Noteborn. 1997. Apoptin induces apoptosis in human transformed and malignant cells but not in normal cells. Proc. Natl. Acad. Sci. USA 94:5843-5847. - PMC - PubMed
-
- Darwich, L., J. Segales, and E. Mateu. 2004. Pathogenesis of postweaning multisystemic wasting syndrome caused by porcine circovirus 2: an immune riddle. Arch. Virol. 149:857-874. [Epub ahead of print.] - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

