The genome of Gryllus bimaculatus nudivirus indicates an ancient diversification of baculovirus-related nonoccluded nudiviruses of insects
- PMID: 17360757
- PMCID: PMC1900193
- DOI: 10.1128/JVI.02781-06
The genome of Gryllus bimaculatus nudivirus indicates an ancient diversification of baculovirus-related nonoccluded nudiviruses of insects
Abstract
The Gryllus bimaculatus nudivirus (GbNV) infects nymphs and adults of the cricket Gryllus bimaculatus (Orthoptera: Gryllidae). GbNV and other nudiviruses such as Heliothis zea nudivirus 1 (HzNV-1) and Oryctes rhinoceros nudivirus (OrNV) were previously called "nonoccluded baculoviruses" as they share some similar structural, genomic, and replication aspects with members of the family Baculoviridae. Their relationships to each other and to baculoviruses are elucidated by the sequence of the complete genome of GbNV, which is 96,944 bp, has an AT content of 72%, and potentially contains 98 predicted protein-coding open reading frames (ORFs). Forty-one ORFs of GbNV share sequence similarities with ORFs found in OrNV, HzNV-1, baculoviruses, and bacteria. Most notably, 15 GbNV ORFs are homologous to the baculovirus core genes, which are associated with transcription (lef-8, lef-9, lef-4, vlf-1, and lef-5), replication (dnapol), structural proteins (p74, pif-1, pif-2, pif-3, vp91, and odv-e56), and proteins of unknown function (38K, ac81, and 19kda). Homologues to these baculovirus core genes have been predicted in HzNV-1 as well. Six GbNV ORFs are homologous to nonconserved baculovirus genes dnaligase, helicase 2, rr1, rr2, iap-3, and desmoplakin. However, the remaining 57 ORFs revealed no homology or poor similarities to the current gene databases. No homologous repeat (hr) sequences but fourteen short direct repeat (dr) regions were detected in the GbNV genome. Gene content and sequence similarity suggest that the nudiviruses GbNV, HzNV-1, and OrNV form a monophyletic group of nonoccluded double-stranded DNA viruses, which separated from the baculovirus lineage before this radiated into dipteran-, hymenopteran-, and lepidopteran-specific clades of occluded nucleopolyhedroviruses and granuloviruses. The accumulated information on the GbNV genome suggests that nudiviruses form a highly diverse and phylogenetically ancient sister group of the baculoviruses, which have evolved in a variety of highly divergent host orders.
Figures
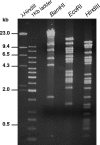



Similar articles
-
Nudiviruses and other large, double-stranded circular DNA viruses of invertebrates: new insights on an old topic.J Invertebr Pathol. 2009 Jul;101(3):187-93. doi: 10.1016/j.jip.2009.03.013. Epub 2009 May 19. J Invertebr Pathol. 2009. PMID: 19460388 Review.
-
The genome of Oryctes rhinoceros nudivirus provides novel insight into the evolution of nuclear arthropod-specific large circular double-stranded DNA viruses.Virus Genes. 2011 Jun;42(3):444-56. doi: 10.1007/s11262-011-0589-5. Epub 2011 Mar 6. Virus Genes. 2011. PMID: 21380757
-
Analysis of the genome of the sexually transmitted insect virus Helicoverpa zea nudivirus 2.Viruses. 2012 Jan;4(1):28-61. doi: 10.3390/v4010028. Epub 2012 Jan 6. Viruses. 2012. PMID: 22355451 Free PMC article.
-
Sequence analysis of a non-classified, non-occluded DNA virus that causes salivary gland hypertrophy of Musca domestica, MdSGHV.Virology. 2008 Jul 20;377(1):184-96. doi: 10.1016/j.virol.2008.04.010. Epub 2008 May 21. Virology. 2008. PMID: 18495197 Free PMC article.
-
The naked truth: An updated review on nudiviruses and their relationship to bracoviruses and baculoviruses.J Invertebr Pathol. 2022 Mar;189:107718. doi: 10.1016/j.jip.2022.107718. Epub 2022 Jan 22. J Invertebr Pathol. 2022. PMID: 35077776 Review.
Cited by
-
MicroRNAs derived from the insect virus HzNV-1 promote lytic infection by suppressing histone methylation.Sci Rep. 2018 Dec 13;8(1):17817. doi: 10.1038/s41598-018-35782-w. Sci Rep. 2018. PMID: 30546025 Free PMC article.
-
A Systematic Review on Viruses in Mass-Reared Edible Insect Species.Viruses. 2021 Nov 15;13(11):2280. doi: 10.3390/v13112280. Viruses. 2021. PMID: 34835086 Free PMC article.
-
Diversity of virophages in metagenomic data sets.J Virol. 2013 Apr;87(8):4225-36. doi: 10.1128/JVI.03398-12. Epub 2013 Feb 13. J Virol. 2013. PMID: 23408616 Free PMC article.
-
Specificity of baculovirus P6.9 basic DNA-binding proteins and critical role of the C terminus in virion formation.J Virol. 2010 Sep;84(17):8821-8. doi: 10.1128/JVI.00072-10. Epub 2010 Jun 2. J Virol. 2010. PMID: 20519380 Free PMC article.
-
The genome and occlusion bodies of marine Penaeus monodon nudivirus (PmNV, also known as MBV and PemoNPV) suggest that it should be assigned to a new nudivirus genus that is distinct from the terrestrial nudiviruses.BMC Genomics. 2014 Jul 25;15(1):628. doi: 10.1186/1471-2164-15-628. BMC Genomics. 2014. PMID: 25063321 Free PMC article.
References
-
- Belloncik, S., and H. Mori. 1998. Cypoviruses. In L. K. Miller and L. A. Ball (ed.), The insect viruses. Plenum Press, New York, NY.
-
- Bideshi, D. K., S. Renault, K. Stasiak, B. A. Federici, and Y. Bigot. 2003. Phylogenetic analysis and possible function of bro-like genes, a multigene family widespread among large double-stranded DNA viruses of invertebrates and bacteria. J. Gen. Virol. 84:2531-2544. - PubMed
-
- Biteau, B., J. Labarre, and M. B. Toledano. 2003. ATP-dependent reduction of cysteine-sulphinic acid by S. cerevisiae sulphiredoxin. Nature 425:980-984. - PubMed
-
- Braunagel, S. C., D. M. Elton, H. Ma, and M. D. Summers. 1996. Identification and analysis of an Autographa californica nuclear polyhedrosis virus structural protein of the occlusion-derived virus envelope: ODV-E56. Virology 217:97-110. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

