Proteomic approach for characterization of hop-inducible proteins in Lactobacillus brevis
- PMID: 17369340
- PMCID: PMC1907096
- DOI: 10.1128/AEM.00124-07
Proteomic approach for characterization of hop-inducible proteins in Lactobacillus brevis
Abstract
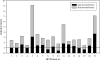
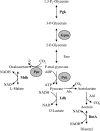
Resistance to hops is a prerequisite for the capability of lactic acid bacteria to grow in beer and thus cause beer spoilage. Bactericidal hop compounds, mainly iso-alpha-acids, are described as ionophores which exchange H+ for cellular divalent cations, e.g., Mn2+, and thus dissipate ion gradients across the cytoplasmic membrane. The acid stress response of Lactobacillus brevis TMW 1.465 and hop adaptation in its variant L. brevis TMW 1.465A caused changes at the level of metabolism, membrane physiology, and cell wall composition. To identify the basis for these changes, a proteomic approach was taken. The experimental design allowed the discrimination of acid stress and hop stress. A strategy for improved protein identification enabled the identification of 84% of the proteins investigated despite the lack of genome sequence data for this strain. Hop resistance in L. brevis TMW 1.465A implies mechanisms to cope with intracellular acidification, mechanisms for energy generation and economy, genetic information fidelity, and enzyme functionality. Interestingly, the majority of hop-regulated enzymes are described as manganese or divalent cation dependent. Regulation of the manganese level allows fine-tuning of the metabolism, which enables a rapid response to environmental (stress) conditions. The hop stress response indicates adaptations shifting the metabolism into an energy-saving mode by effective substrate conversion and prevention of exhaustive protein de novo synthesis. The findings further demonstrate that hop stress in bacteria not only is associated with proton motive force depletion but obviously implies divalent cation limitation.
Figures


Similar articles
-
Characterization of a highly hop-resistant Lactobacillus brevis strain lacking hop transport.Appl Environ Microbiol. 2006 Oct;72(10):6483-92. doi: 10.1128/AEM.00668-06. Appl Environ Microbiol. 2006. PMID: 17021196 Free PMC article.
-
Detection of acid and hop shock induced responses in beer spoiling Lactobacillus brevis by MALDI-TOF MS.Food Microbiol. 2015 Apr;46:501-506. doi: 10.1016/j.fm.2014.09.018. Epub 2014 Oct 7. Food Microbiol. 2015. PMID: 25475321
-
Mechanisms of hop inhibition: hop ionophores.J Agric Food Chem. 2009 Jul 22;57(14):6074-81. doi: 10.1021/jf900847y. J Agric Food Chem. 2009. PMID: 19537790
-
Beer spoilage bacteria and hop resistance.Int J Food Microbiol. 2003 Dec 31;89(2-3):105-24. doi: 10.1016/s0168-1605(03)00153-3. Int J Food Microbiol. 2003. PMID: 14623377 Review.
-
Differential proteomic analysis in the study of prokaryotes stress resistance.Ann Ist Super Sanita. 2005;41(4):459-68. Ann Ist Super Sanita. 2005. PMID: 16569914 Review.
Cited by
-
Cell Envelope Modifications Generating Resistance to Hop Beta Acids and Collateral Sensitivity to Cationic Antimicrobials in Listeria monocytogenes.Microorganisms. 2023 Aug 7;11(8):2024. doi: 10.3390/microorganisms11082024. Microorganisms. 2023. PMID: 37630584 Free PMC article.
-
MALDI-TOF Mass Spectrometry Enables a Comprehensive and Fast Analysis of Dynamics and Qualities of Stress Responses of Lactobacillus paracasei subsp. paracasei F19.PLoS One. 2016 Oct 26;11(10):e0165504. doi: 10.1371/journal.pone.0165504. eCollection 2016. PLoS One. 2016. PMID: 27783652 Free PMC article.
-
Characterization of the Adaptive Amoxicillin Resistance of Lactobacillus casei Zhang by Proteomic Analysis.Front Microbiol. 2018 Feb 20;9:292. doi: 10.3389/fmicb.2018.00292. eCollection 2018. Front Microbiol. 2018. PMID: 29515561 Free PMC article.
-
Gene expression of lactobacilli in murine forestomach biofilms.Microb Biotechnol. 2014 Jul;7(4):347-59. doi: 10.1111/1751-7915.12126. Epub 2014 Apr 4. Microb Biotechnol. 2014. PMID: 24702817 Free PMC article.
-
Transcriptome Analysis of Viable but Non-Culturable Brettanomyces bruxellensis Induced by Hop Bitter Acids.Front Microbiol. 2022 May 30;13:902110. doi: 10.3389/fmicb.2022.902110. eCollection 2022. Front Microbiol. 2022. PMID: 35707174 Free PMC article.
References
-
- Albrechtsen, H., and S. I. Ahmad. 1980. Regulation of the synthesis of nucleoside catabolic enzymes in Escherichia coli: further analysis of a deo Oc mutant strain. Mol. Gen. Genet. 179:457-460. - PubMed
-
- Alonso, J. C., A. C. Stiege, B. Dobrinski, and R. Lurz. 1993. Purification and properties of the RecR protein from Bacillus subtilis 168. J. Biol. Chem. 268:1424-1429. - PubMed
-
- Back, W. 1994. Farbatlas und Handbuch der Getränkebiologie. Verlag Hans Carl, Nürnberg, Germany.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

