Ultraconserved elements are associated with homeostatic control of splicing regulators by alternative splicing and nonsense-mediated decay
- PMID: 17369403
- PMCID: PMC1820944
- DOI: 10.1101/gad.1525507
Ultraconserved elements are associated with homeostatic control of splicing regulators by alternative splicing and nonsense-mediated decay
Abstract
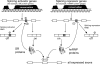
Many alternative splicing events create RNAs with premature stop codons, suggesting that alternative splicing coupled with nonsense-mediated decay (AS-NMD) may regulate gene expression post-transcriptionally. We tested this idea in mice by blocking NMD and measuring changes in isoform representation using splicing-sensitive microarrays. We found a striking class of highly conserved stop codon-containing exons whose inclusion renders the transcript sensitive to NMD. A genomic search for additional examples identified>50 such exons in genes with a variety of functions. These exons are unusually frequent in genes that encode splicing activators and are unexpectedly enriched in the so-called "ultraconserved" elements in the mammalian lineage. Further analysis show that NMD of mRNAs for splicing activators such as SR proteins is triggered by splicing activation events, whereas NMD of the mRNAs for negatively acting hnRNP proteins is triggered by splicing repression, a polarity consistent with widespread homeostatic control of splicing regulator gene expression. We suggest that the extreme genomic conservation surrounding these regulatory splicing events within splicing factor genes demonstrates the evolutionary importance of maintaining tightly tuned homeostasis of RNA-binding protein levels in the vertebrate cell.
Figures
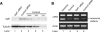


References
-
- Ashburner M., Ball C.A., Blake J.A., Botstein D., Butler H., Cherry J.M., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Ball C.A., Blake J.A., Botstein D., Butler H., Cherry J.M., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Blake J.A., Botstein D., Butler H., Cherry J.M., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Botstein D., Butler H., Cherry J.M., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Butler H., Cherry J.M., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Cherry J.M., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Dolinski K., Dwight S.S., Eppig J.T., Dwight S.S., Eppig J.T., Eppig J.T., et al. Gene ontology: Tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 2000;25:25–29. - PMC - PubMed
-
- Bejerano G., Pheasant M., Makunin I., Stephen S., Kent W.J., Mattick J.S., Haussler D., Pheasant M., Makunin I., Stephen S., Kent W.J., Mattick J.S., Haussler D., Makunin I., Stephen S., Kent W.J., Mattick J.S., Haussler D., Stephen S., Kent W.J., Mattick J.S., Haussler D., Kent W.J., Mattick J.S., Haussler D., Mattick J.S., Haussler D., Haussler D. Ultraconserved elements in the human genome. Science. 2004;304:1321–1325. - PubMed
-
- Bejerano G., Lowe C.B., Ahituv N., King B., Siepel A., Salama S.R., Rubin E.M., Kent W.J., Haussler D., Lowe C.B., Ahituv N., King B., Siepel A., Salama S.R., Rubin E.M., Kent W.J., Haussler D., Ahituv N., King B., Siepel A., Salama S.R., Rubin E.M., Kent W.J., Haussler D., King B., Siepel A., Salama S.R., Rubin E.M., Kent W.J., Haussler D., Siepel A., Salama S.R., Rubin E.M., Kent W.J., Haussler D., Salama S.R., Rubin E.M., Kent W.J., Haussler D., Rubin E.M., Kent W.J., Haussler D., Kent W.J., Haussler D., Haussler D. A distal enhancer and an ultraconserved exon are derived from a novel retroposon. Nature. 2006;441:87–90. - PubMed
-
- Bell L.R., Horabin J.I., Schedl P., Cline T.W., Horabin J.I., Schedl P., Cline T.W., Schedl P., Cline T.W., Cline T.W. Positive autoregulation of sex-lethal by alternative splicing maintains the female determined state in Drosophila. Cell. 1991;65:229–239. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
