Genomotyping of Pseudomonas putida strains using P. putida KT2440-based high-density DNA microarrays: implications for transcriptomics studies
- PMID: 17370070
- PMCID: PMC1914237
- DOI: 10.1007/s00253-007-0914-z
Genomotyping of Pseudomonas putida strains using P. putida KT2440-based high-density DNA microarrays: implications for transcriptomics studies
Abstract
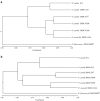
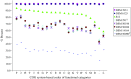
Pseudomonas putida KT2440 is the only fully sequenced P. putida strain. Thus, for transcriptomics and proteomics studies with other P. putida strains, the P. putida KT2440 genomic database serves as standard reference. The utility of KT2440 whole-genome, high-density oligonucleotide microarrays for transcriptomics studies of other Pseudomonas strains was investigated. To this end, microarray hybridizations were performed with genomic DNAs of subcultures of P. putida KT2440 (DSM6125), the type strain (DSM291(T)), plasmid pWW0-containing KT2440-derivative strain mt-2 (DSM3931), the solvent-tolerant P. putida S12, and several other Pseudomonas strains. Depending on the strain tested, 22 to 99% of all genetic elements were identified in the genomic DNAs. The efficacy of these microarrays to study cellular function was determined for all strains included in the study. The vast majority of DSM6125 genes encoding proteins of primary metabolism and genes involved in the catabolism of aromatic compounds were identified in the genomic DNA of strain S12: a prerequisite for reliable transcriptomics analyses. The genomotypic comparisons between Pseudomonas strains were used to construct highly discriminative phylogenetic relationships. DSM6125 and DSM3931 were indistinguishable and clustered together with strain S12 in a separate group, distinct from DSM291(T). Pseudomonas monteilii (DSM14164) clustered well with P. putida strains.
Figures
References
-
- Brosch R, Lefevre M, Grimont F, Grimont PAD. Taxonomic diversity of pseudomonads revealed by computer interpretation of ribotyping data. Syst Appl Microbiol. 1996;19:541–555.
-
- Dabboussi F, Hamze M, Singer E, Geoffroy V, Meyer JM, Izard D. Pseudomonas mosselii sp. nov., a novel species isolated from clinical specimens. Int J Syst Evol Microbiol. 2002;52:363–376. - PubMed
-
- De Bont JAM. Solvent-tolerant bacteria in biocatalysis. Trends Biotechnol. 1998;16:493–499.
-
- Dominguez-Cuevas P, Gonzalez-Pastor JE, Marques S, Ramos JL, de Lorenzo V. Transcriptional tradeoff between metabolic and stress-response programs in Pseudomonas putida KT2440 cells exposed to toluene. J Biol Chem. 2006;281:11981–11991. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

