Subtypes of Cryptosporidium parvum in humans and disease risk
- PMID: 17370519
- PMCID: PMC2725800
- DOI: 10.3201/eid1301.060481
Subtypes of Cryptosporidium parvum in humans and disease risk
Abstract
The 2 main species of Cryptosporidium that infect humans are Cryptosporidium hominis and C. parvum. Here, multilocus fragment analysis of 3 microsatellite loci (ML1, ML2, and gp60) was used to subtype strains from sporadic cases of cryptosporidiosis in Wales and northwest England. Of 72 strains of C. parvum, 63 were typeable at all 3 loci, forming 31 subtypes. These strains formed 3 broad clusters, representing 74.6%, 20.6%, and 4.8% of typeable strains. Of 118 C. hominis strains, 106 were typeable at all 3 loci, forming 9 subtypes; however, 90% belonged to the same subtype. Analysis with epidemiologic data found an association between strains from case-patients who reported contact with farm animals and individual C. parvum microsatellite alleles. The strongest association was with ML1; all strains from case-patients that reported farm animal contact had the same allele (ML1-242). Microsatellite typing of C. parvum provides valuable additional information on the epidemiology of this pathogen.
Figures
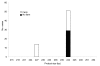
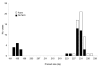
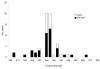
References
-
- McLauchlin J, Amar C, Pedraza-Díaz S, Nichols GL. Molecular epidemiological analysis of Cryptosporidium spp. in the United Kingdom: results of genotyping Cryptosporidium spp. in 1705 fecal samples from humans and 105 fecal samples from livestock animals. J Clin Microbiol. 2000;38:3984–90. - PMC - PubMed
-
- The development of a national collection for oocysts of Cryptosporidium. Foundation for Water Research, Marlow, Bucks, UK, 2002. [cited 2006 Oct 11]. Available from http://www.fwr.org/
-
- Health Protection Agency. 2006. [cited 2006 Oct 11]. Available from http://www.hpa.org.uk/
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
