Insights on the role of nucleic acid/protein interactions in chaperoned nucleic acid rearrangements of HIV-1 reverse transcription
- PMID: 17372205
- PMCID: PMC1828707
- DOI: 10.1073/pnas.0700166104
Insights on the role of nucleic acid/protein interactions in chaperoned nucleic acid rearrangements of HIV-1 reverse transcription
Abstract
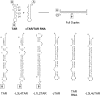
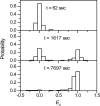
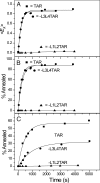
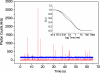
HIV-1 reverse transcription requires several nucleic acid rearrangement steps that are "chaperoned" by the nucleocapsid protein (NC), including minus-strand transfer, in which the DNA transactivation response element (TAR) is annealed to the complementary TAR RNA region of the viral genome. These various rearrangement processes occur in NC bound complexes of specific RNA and DNA structures. A major barrier to the investigation of these processes in vitro has been the diversity and heterogeneity of the observed nucleic acid/protein assemblies, ranging from small complexes of only one or two nucleic acid molecules all the way up to large-scale aggregates comprised of thousands of NC and nucleic acid molecules. Herein, we use a flow chamber approach involving rapid NC/nucleic acid mixing to substantially control aggregation for the NC chaperoned irreversible annealing kinetics of a model TAR DNA hairpin sequence to the complementary TAR RNA hairpin, i.e., to form an extended duplex. By combining the flow chamber approach with a broad array of fluorescence single-molecule spectroscopy (SMS) tools (FRET, molecule counting, and correlation spectroscopy), we have unraveled the complex, heterogeneous kinetics that occur during the course of annealing. The SMS results demonstrate that the TAR hairpin reactant is predominantly a single hairpin coated by multiple NCs with a dynamic secondary structure, involving equilibrium between a "Y" shaped conformation and a closed one. The data further indicate that the nucleation of annealing occurs in an encounter complex that is formed by two hairpins with one or both of the hairpins in the "Y" conformation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

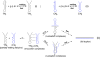
Similar articles
-
Single-molecule FRET studies of important intermediates in the nucleocapsid-protein-chaperoned minus-strand transfer step in HIV-1 reverse transcription.Biophys J. 2005 Nov;89(5):3470-9. doi: 10.1529/biophysj.105.065326. Epub 2005 Aug 12. Biophys J. 2005. PMID: 16100256 Free PMC article.
-
Probing nucleation, reverse annealing, and chaperone function along the reaction path of HIV-1 single-strand transfer.Proc Natl Acad Sci U S A. 2007 Jul 31;104(31):12651-6. doi: 10.1073/pnas.0700350104. Epub 2007 Jun 19. Proc Natl Acad Sci U S A. 2007. PMID: 17578926 Free PMC article.
-
Mechanistic studies of mini-TAR RNA/DNA annealing in the absence and presence of HIV-1 nucleocapsid protein.J Mol Biol. 2006 Oct 13;363(1):244-61. doi: 10.1016/j.jmb.2006.08.039. Epub 2006 Aug 22. J Mol Biol. 2006. PMID: 16962137
-
Initiation of HIV-1 reverse transcription and functional role of nucleocapsid-mediated tRNA/viral genome interactions.Virus Res. 2012 Nov;169(2):324-39. doi: 10.1016/j.virusres.2012.06.006. Epub 2012 Jun 18. Virus Res. 2012. PMID: 22721779 Review.
-
Nucleic acid chaperone activity of HIV-1 nucleocapsid protein: critical role in reverse transcription and molecular mechanism.Prog Nucleic Acid Res Mol Biol. 2005;80:217-86. doi: 10.1016/S0079-6603(05)80006-6. Prog Nucleic Acid Res Mol Biol. 2005. PMID: 16164976 Review. No abstract available.
Cited by
-
The nucleic acid chaperone activity of the HIV-1 Gag polyprotein is boosted by its cellular partner RPL7: a kinetic study.Nucleic Acids Res. 2020 Sep 18;48(16):9218-9234. doi: 10.1093/nar/gkaa659. Nucleic Acids Res. 2020. PMID: 32797159 Free PMC article.
-
Structural Insights into the HIV-1 Minus-strand Strong-stop DNA.J Biol Chem. 2016 Feb 12;291(7):3468-82. doi: 10.1074/jbc.M115.708099. Epub 2015 Dec 14. J Biol Chem. 2016. PMID: 26668324 Free PMC article.
-
Comparative analysis of RNA/protein dynamics for the arginine-rich-binding motif and zinc-finger-binding motif proteins encoded by HIV-1.Biophys J. 2010 Nov 17;99(10):3454-62. doi: 10.1016/j.bpj.2010.09.051. Biophys J. 2010. PMID: 21081095 Free PMC article.
-
Single-molecule spectroscopic study of dynamic nanoscale DNA bending behavior of HIV-1 nucleocapsid protein.J Phys Chem B. 2013 Apr 25;117(16):4183-96. doi: 10.1021/jp3018259. Epub 2012 May 16. J Phys Chem B. 2013. PMID: 22591315 Free PMC article.
-
Nucleocapsid Protein: A Desirable Target for Future Therapies Against HIV-1.Curr Top Microbiol Immunol. 2015;389:53-92. doi: 10.1007/82_2015_433. Curr Top Microbiol Immunol. 2015. PMID: 25749978 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

