Infectivity determinants of the hepatitis B virus pre-S domain are confined to the N-terminal 75 amino acid residues
- PMID: 17376925
- PMCID: PMC1900317
- DOI: 10.1128/JVI.00096-07
Infectivity determinants of the hepatitis B virus pre-S domain are confined to the N-terminal 75 amino acid residues
Abstract
The N-terminal pre-S domain of the large hepatitis B virus (HBV) envelope protein plays a pivotal role at the initial step of the viral entry pathway. In the present study, the entire pre-S domain was mapped for infectivity determinants, following a reverse-genetics approach and using in vitro infection assays with hepatitis delta virus (HDV) or HBV particles. The results demonstrate that lesions created within the N-terminal 75 amino acids of the pre-S region abrogate infectivity, whereas mutations between amino acids 76 and 113, overlapping the matrix domain, had no effect. In contrast to the results of a recent study (L. Stoeckl, A. Funk, A. Kopitzki, B. Brandenburg, S. Oess, H. Will, H. Sirma, and E. Hildt, Proc. Natl. Acad. Sci. 103:6730-6734, 2006), the deletion of a cell membrane translocation motif (TLM) located between amino acids 148 and 161 at the C terminus of pre-S2 did not interfere with the infectivity of the resulting HDV or HBV mutants. Furthermore, a series of large deletions overlapping the pre-S2 domain were compatible with infectivity, although the efficiency of infection was reduced when the deletions extended to the pre-S1 domain. Overall, the results demonstrate that the activity of the pre-S domain at viral entry solely depends on the integrity of its first 75 amino acids and thus excludes any function of the matrix domain or TLM.
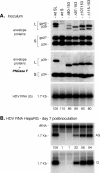
Figures







References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

