Synthesis and function of membrane phosphoinositides in budding yeast, Saccharomyces cerevisiae
- PMID: 17382260
- PMCID: PMC1868553
- DOI: 10.1016/j.bbalip.2007.01.015
Synthesis and function of membrane phosphoinositides in budding yeast, Saccharomyces cerevisiae
Abstract
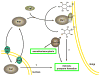
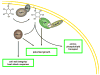
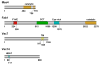
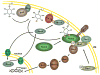
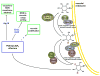
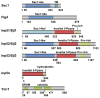
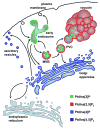
It is now well appreciated that derivatives of phosphatidylinositol (PtdIns) are key regulators of many cellular processes in eukaryotes. Of particular interest are phosphoinositides (mono- and polyphosphorylated adducts to the inositol ring in PtdIns), which are located at the cytoplasmic face of cellular membranes. Phosphoinositides serve both a structural and a signaling role via their recruitment of proteins that contain phosphoinositide-binding domains. Phosphoinositides also have a role as precursors of several types of second messengers for certain intracellular signaling pathways. Realization of the importance of phosphoinositides has brought increased attention to characterization of the enzymes that regulate their synthesis, interconversion, and turnover. Here we review the current state of our knowledge about the properties and regulation of the ATP-dependent lipid kinases responsible for synthesis of phosphoinositides and also the additional temporal and spatial controls exerted by the phosphatases and a phospholipase that act on phosphoinositides in yeast.
Figures

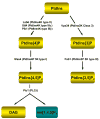

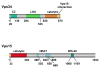

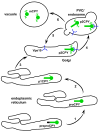

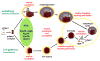

References
-
- Carpenter CL, Cantley LC. Phosphoinositide kinases. Current Opinion in Cell Biology. 1996;8:153–8. - PubMed
-
- Fruman DA, Meyers RE, Cantley LC. Phosphoinositide kinases. Annual Review of Biochemistry. 1998;67:481–507. - PubMed
-
- Majerus PW. Inositol phosphatases and kinases in cell signaling. Faseb Journal. 1997;11:A1297–A1297.
-
- Majerus PW, Kisseleva MV, Norris FA. The role of phosphatases in inositol signaling reactions. Journal of Biological Chemistry. 1999;274:10669–10672. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

