Mutations in rsmG, encoding a 16S rRNA methyltransferase, result in low-level streptomycin resistance and antibiotic overproduction in Streptomyces coelicolor A3(2)
- PMID: 17384192
- PMCID: PMC1913335
- DOI: 10.1128/JB.01776-06
Mutations in rsmG, encoding a 16S rRNA methyltransferase, result in low-level streptomycin resistance and antibiotic overproduction in Streptomyces coelicolor A3(2)
Abstract
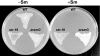
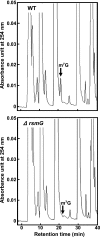
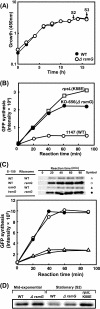
Certain str mutations that confer high- or low-level streptomycin resistance result in the overproduction of antibiotics by Streptomyces spp. The str mutations that confer the high-level resistance occur within rpsL, which encodes the ribosomal protein S12, while those that cause low-level resistance are not as well known. We have used comparative genome sequencing to determine that low-level resistance is caused by mutations of rsmG, which encodes an S-adenosylmethionine (SAM)-dependent 16S rRNA methyltransferase containing a SAM binding motif. Deletion of rsmG from wild-type Streptomyces coelicolor resulted in the acquisition of streptomycin resistance and the overproduction of the antibiotic actinorhodin. Introduction of wild-type rsmG into the deletion mutant completely abrogated the effects of the rsmG deletion, confirming that rsmG mutation underlies the observed phenotype. Consistent with earlier work using a spontaneous rsmG mutant, the strain carrying DeltarsmG exhibited increased SAM synthetase activity, which mediated the overproduction of antibiotic. Moreover, high-performance liquid chromatography analysis showed that the DeltarsmG mutant lacked a 7-methylguanosine modification in the 16S rRNA (possibly at position G518, which corresponds to G527 of Escherichia coli). Like certain rpsL mutants, the DeltarsmG mutant exhibited enhanced protein synthetic activity during the late growth phase. Unlike rpsL mutants, however, the DeltarsmG mutant showed neither greater stability of the 70S ribosomal complex nor increased expression of ribosome recycling factor, suggesting that the mechanism underlying increased protein synthesis differs in the rsmG and the rpsL mutants. Finally, spontaneous rsmG mutations arose at a 1,000-fold-higher frequency than rpsL mutations. These findings provide new insight into the role of rRNA modification in activating secondary metabolism in Streptomyces.
Figures


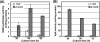
Comment in
-
Novel links between antibiotic resistance and antibiotic production.J Bacteriol. 2007 May;189(10):3683-5. doi: 10.1128/JB.00356-07. Epub 2007 Mar 23. J Bacteriol. 2007. PMID: 17384190 Free PMC article. Review. No abstract available.
Similar articles
-
Identification of the RsmG methyltransferase target as 16S rRNA nucleotide G527 and characterization of Bacillus subtilis rsmG mutants.J Bacteriol. 2007 Aug;189(16):6068-73. doi: 10.1128/JB.00558-07. Epub 2007 Jun 15. J Bacteriol. 2007. PMID: 17573471 Free PMC article.
-
Impairing methylations at ribosome RNA, a point mutation-dependent strategy for aminoglycoside resistance: the rsmG case.Biomedica. 2014 Apr;34 Suppl 1:41-9. doi: 10.1590/S0120-41572014000500006. Biomedica. 2014. PMID: 24968035
-
Loss of a conserved 7-methylguanosine modification in 16S rRNA confers low-level streptomycin resistance in bacteria.Mol Microbiol. 2007 Feb;63(4):1096-106. doi: 10.1111/j.1365-2958.2006.05585.x. Mol Microbiol. 2007. PMID: 17238915
-
Novel links between antibiotic resistance and antibiotic production.J Bacteriol. 2007 May;189(10):3683-5. doi: 10.1128/JB.00356-07. Epub 2007 Mar 23. J Bacteriol. 2007. PMID: 17384190 Free PMC article. Review. No abstract available.
-
Combinatorial genetic technology for the development of new anti-infectives.Arch Pathol Lab Med. 2004 Dec;128(12):1351-9. doi: 10.5858/2004-128-1351-CGTFTD. Arch Pathol Lab Med. 2004. PMID: 15578878 Review.
Cited by
-
Insights into microbial cryptic gene activation and strain improvement: principle, application and technical aspects.J Antibiot (Tokyo). 2017 Jan;70(1):25-40. doi: 10.1038/ja.2016.82. Epub 2016 Jul 6. J Antibiot (Tokyo). 2017. PMID: 27381522 Review.
-
Identification of the RsmG methyltransferase target as 16S rRNA nucleotide G527 and characterization of Bacillus subtilis rsmG mutants.J Bacteriol. 2007 Aug;189(16):6068-73. doi: 10.1128/JB.00558-07. Epub 2007 Jun 15. J Bacteriol. 2007. PMID: 17573471 Free PMC article.
-
Ribosome biogenesis and the translation process in Escherichia coli.Microbiol Mol Biol Rev. 2007 Sep;71(3):477-94. doi: 10.1128/MMBR.00013-07. Microbiol Mol Biol Rev. 2007. PMID: 17804668 Free PMC article. Review.
-
Efficiently activated ε-poly-L-lysine production by multiple antibiotic-resistance mutations and acidic pH shock optimization in Streptomyces albulus.Microbiologyopen. 2019 May;8(5):e00728. doi: 10.1002/mbo3.728. Epub 2018 Oct 8. Microbiologyopen. 2019. PMID: 30298553 Free PMC article.
-
Activation and products of the cryptic secondary metabolite biosynthetic gene clusters by rifampin resistance (rpoB) mutations in actinomycetes.J Bacteriol. 2013 Jul;195(13):2959-70. doi: 10.1128/JB.00147-13. Epub 2013 Apr 19. J Bacteriol. 2013. PMID: 23603745 Free PMC article.
References
-
- Albert, T. J., D. Dailidiene, G. Dailide, J. E. Norton, A. Kalia, T. A. Richmond, M. Molla, J. Singh, R. D. Green, and D. E. Berg. 2005. Mutation discovery in bacterial genomes: metronidazole resistance in Helicobacter pylori. Nat. Methods 2:951-953. - PubMed
-
- Andersson, D. I., and B. R. Levin. 1999. The biological cost of antibiotic resistance. Curr. Opin. Microbiol. 2:489-493. - PubMed
-
- Bibb, M. J. 2005. Regulation of secondary metabolism in streptomycetes. Curr. Opin. Microbiol. 8:208-215. - PubMed
-
- Carter, A. P., W. M. Clemons, D. E. Brodersen, R. J. Morgan-Warren, B. T. Wimberly, and V. Ramakrishnan. 2000. Functional insights from the structure of the 30S ribosomal subunit and its interactions with antibiotics. Nature 407:340-348. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

