Characterizing linkage disequilibrium in pig populations
- PMID: 17384735
- PMCID: PMC1802018
- DOI: 10.7150/ijbs.3.166
Characterizing linkage disequilibrium in pig populations
Abstract
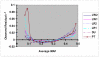
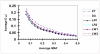
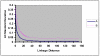
Knowledge of the extent and range of linkage disequilibrium (LD), defined as non-random association of alleles at two or more loci, in animal populations is extremely valuable in localizing genes affecting quantitative traits, identifying chromosomal regions under selection, studying population history, and characterizing/managing genetic resources and diversity. Two commonly used LD measures, r(2) and D', and their permutation based adjustments, were evaluated using genotypes of more than 6,000 pigs from six commercial lines (two terminal sire lines and four maternal lines) at ~4,500 autosomal SNPs (single nucleotide polymorphisms). The results indicated that permutation only partially removed the dependency of D' on allele frequency and that r(2) is a considerably more robust LD measure. The maximum r(2) was derived as a function of allele frequency. Using the same genotype dataset, the extent of LD in these pig populations was estimated for all possible syntenic SNP pairs using r(2) and the ratio of r(2) over its theoretical maximum. As expected, the extent of LD highest for SNP pairs was found in tightest linkage and decreased as their map distance increased. The level of LD found in these pig populations appears to be lower than previously implied in several other studies using microsatellite genotype data. For all pairs of SNPs approximately 3 centiMorgan (cM) apart, the average r(2) was equal to 0.1. Based on the average population-wise LD found in these six commercial pig lines, we recommend a spacing of 0.1 to 1 cM for a whole genome association study in pig populations.
Conflict of interest statement
Conflict of interest: The authors have declared that no conflict of interest exists.
Figures






References
-
- Hartl DL, Clark AG. Principle of Population Genetics. Sinauer Associates. 1997.
-
- Bulmer MG. The effect of selection on genetic variability. The American Naturalist. 1971;105:201–211.
-
- Sved JA. Linkage disequilibrium and homozygosity of chromosome segments in finite populations. Theoretical Population Biology. 1971;2:125–141. - PubMed
-
- Ardlie KG, Kruglyak L, Seielstad M. Patterns of linkage disequilibrium in the human genom. Nat. Rev. Genet. 2002;4:299–309. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

