Stable complexes formed by HIV-1 reverse transcriptase at distinct positions on the primer-template controlled by binding deoxynucleoside triphosphates or foscarnet
- PMID: 17400246
- PMCID: PMC1986715
- DOI: 10.1016/j.jmb.2007.03.006
Stable complexes formed by HIV-1 reverse transcriptase at distinct positions on the primer-template controlled by binding deoxynucleoside triphosphates or foscarnet
Abstract
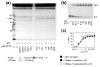
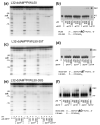
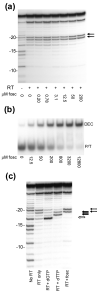
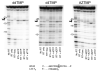
Binding of the next complementary dNTP by the binary complex containing HIV-1 reverse transcriptase (RT) and primer-template induces conformational changes that have been implicated in catalytic function of RT. We have used DNase I footprinting, gel electrophoretic mobility shift, and exonuclease protection assays to characterize the interactions between HIV-1 RT and chain-terminated primer-template in the absence and presence of various ligands. Distinguishable stable complexes were formed in the presence of foscarnet (an analog of pyrophosphate), the dNTP complementary to the first (+1) templating nucleotide or the dNTP complementary to the second (+2) templating nucleotide. The position of HIV-1 RT on the primer-template in each of these complexes is different. RT is located upstream in the foscarnet complex, relative to the +1 complex, and downstream in the +2 complex. These results suggest that HIV-1 RT can translocate along the primer-template in the absence of phosphodiester bond formation. The ability to form a specific foscarnet complex might explain the inhibitory properties of this compound. The ability to recognize the second templating nucleotide has implications for nucleotide misincorporation.
Figures



Similar articles
-
Interactions between HIV-1 reverse transcriptase and the downstream template strand in stable complexes with primer-template.PLoS One. 2008;3(10):e3561. doi: 10.1371/journal.pone.0003561. Epub 2008 Oct 30. PLoS One. 2008. PMID: 18974785 Free PMC article.
-
Relationship between 3'-azido-3'-deoxythymidine resistance and primer unblocking activity in foscarnet-resistant mutants of human immunodeficiency virus type 1 reverse transcriptase.J Virol. 2003 Jun;77(11):6127-37. doi: 10.1128/jvi.77.11.6127-6137.2003. J Virol. 2003. PMID: 12743270 Free PMC article.
-
Nucleotide-induced stable complex formation by HIV-1 reverse transcriptase.Biochemistry. 1997 May 13;36(19):5749-57. doi: 10.1021/bi962410z. Biochemistry. 1997. PMID: 9153415
-
Protein-nucleic acid interactions and DNA conformation in a complex of human immunodeficiency virus type 1 reverse transcriptase with a double-stranded DNA template-primer.Biopolymers. 1997;44(2):125-38. doi: 10.1002/(SICI)1097-0282(1997)44:2<125::AID-BIP2>3.0.CO;2-X. Biopolymers. 1997. PMID: 9354757 Review.
-
Effects of nucleotides and nucleotide analogue inhibitors of HIV-1 reverse transcriptase in a ratchet model of polymerase translocation.Curr Pharm Des. 2006;12(15):1867-77. doi: 10.2174/138161206776873626. Curr Pharm Des. 2006. PMID: 16724953 Review.
Cited by
-
Interactions between HIV-1 reverse transcriptase and the downstream template strand in stable complexes with primer-template.PLoS One. 2008;3(10):e3561. doi: 10.1371/journal.pone.0003561. Epub 2008 Oct 30. PLoS One. 2008. PMID: 18974785 Free PMC article.
-
Structures of reverse transcriptase pre- and post-excision complexes shed new light on HIV-1 AZT resistance.Viruses. 2011 Jan;3(1):20-25. doi: 10.3390/v3010020. Epub 2011 Jan 18. Viruses. 2011. PMID: 21980583 Free PMC article.
-
Connection domain mutations N348I and A360V in HIV-1 reverse transcriptase enhance resistance to 3'-azido-3'-deoxythymidine through both RNase H-dependent and -independent mechanisms.J Biol Chem. 2008 Aug 8;283(32):22222-32. doi: 10.1074/jbc.M803521200. Epub 2008 Jun 10. J Biol Chem. 2008. PMID: 18547911 Free PMC article.
-
Exploring the role of the α-carboxyphosphonate moiety in the HIV-RT activity of α-carboxy nucleoside phosphonates.Org Biomol Chem. 2016 Feb 28;14(8):2454-65. doi: 10.1039/c5ob02507a. Org Biomol Chem. 2016. PMID: 26813581 Free PMC article.
-
Small Molecule Drugs Targeting Viral Polymerases.Pharmaceuticals (Basel). 2024 May 20;17(5):661. doi: 10.3390/ph17050661. Pharmaceuticals (Basel). 2024. PMID: 38794231 Free PMC article. Review.
References
-
- Johnson KA. Conformational coupling in DNA polymerase fidelity. Annu. Rev. Biochem. 1993;62:685–713. - PubMed
-
- Landick R. Active-site dynamics in RNA polymerases. Cell. 2004;116:351–353. - PubMed
-
- Mizrahi V, Henrie RN, Marlier JF, Johnson KA, Benkovic SJ. Rate-limiting steps in the DNA polymerase I reaction pathway. Biochemistry. 1985;24:4010–4018. - PubMed
-
- Kuchta RD, Mizrahi V, Benkovic PA, Johnson KA, Benkovic SJ. Kinetic mechanism of DNA polymerase I (Klenow) Biochemistry. 1987;26:8410–8417. - PubMed
-
- Patel SS, Wong I, Johnson KA. Pre-steady-state kinetic analysis of processive DNA replication including complete characterization of an exonuclease-deficient mutant. Biochemistry. 1991;30:511–525. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

