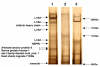Identification of potential protein interactors of Lrrk2
- PMID: 17400507
- PMCID: PMC2970619
- DOI: 10.1016/j.parkreldis.2007.01.008
Identification of potential protein interactors of Lrrk2
Abstract
Pathogenic substitutions in the Lrrk2 protein have been shown to be an important cause of both familial and sporadic parkinsonism. The molecular pathway involved in Lrrk2 dopaminergic neuron degeneration remains elusive. Employing a combination of Lrrk2-mediated protein precipitation and tandem mass spectrometry, we identified 14 potential Lrrk2 binding partners. The majority of these interactions may be subgrouped into three functional cellular pathways: (i) chaperone-mediated response, (ii) proteins associated with the cytoskeleton and trafficking and (iii) phosphorylation and kinase activity. Future investigation of these candidates is now warranted and may help resolve the pathomechanism behind Lrrk2 neurodegeneration.
Figures
Similar articles
-
The chaperone activity of heat shock protein 90 is critical for maintaining the stability of leucine-rich repeat kinase 2.J Neurosci. 2008 Mar 26;28(13):3384-91. doi: 10.1523/JNEUROSCI.0185-08.2008. J Neurosci. 2008. PMID: 18367605 Free PMC article.
-
A QUICK screen for Lrrk2 interaction partners--leucine-rich repeat kinase 2 is involved in actin cytoskeleton dynamics.Mol Cell Proteomics. 2011 Jan;10(1):M110.001172. doi: 10.1074/mcp.M110.001172. Epub 2010 Sep 27. Mol Cell Proteomics. 2011. PMID: 20876399 Free PMC article.
-
A comparative analysis of leucine-rich repeat kinase 2 (Lrrk2) expression in mouse brain and Lewy body disease.Neuroscience. 2007 Jul 29;147(4):1047-58. doi: 10.1016/j.neuroscience.2007.05.027. Epub 2007 Jul 3. Neuroscience. 2007. PMID: 17611037
-
On the road to leucine-rich repeat kinase 2 signalling: evidence from cellular and in vivo studies.Neurosignals. 2011;19(1):1-15. doi: 10.1159/000324488. Epub 2011 Mar 23. Neurosignals. 2011. PMID: 21430363 Review.
-
Phosphorylation of LRRK2: from kinase to substrate.Biochem Soc Trans. 2012 Oct;40(5):1102-10. doi: 10.1042/BST20120128. Biochem Soc Trans. 2012. PMID: 22988873 Review.
Cited by
-
Regulation of LRRK2 stability by the E3 ubiquitin ligase CHIP.PLoS One. 2009 Jun 17;4(6):e5949. doi: 10.1371/journal.pone.0005949. PLoS One. 2009. PMID: 19536328 Free PMC article.
-
The amyloid precursor protein/protease nexin 2 Kunitz inhibitor domain is a highly specific substrate of mesotrypsin.J Biol Chem. 2010 Jan 15;285(3):1939-49. doi: 10.1074/jbc.M109.057216. Epub 2009 Nov 17. J Biol Chem. 2010. PMID: 19920152 Free PMC article.
-
LRRK2 Pathways Leading to Neurodegeneration.Curr Neurol Neurosci Rep. 2015 Jul;15(7):42. doi: 10.1007/s11910-015-0564-y. Curr Neurol Neurosci Rep. 2015. PMID: 26008812 Free PMC article. Review.
-
Purine-scaffold Hsp90 inhibitors.Curr Top Med Chem. 2009;9(15):1436-46. doi: 10.2174/156802609789895737. Curr Top Med Chem. 2009. PMID: 19860732 Free PMC article. Review.
-
Cellular processes associated with LRRK2 function and dysfunction.FEBS J. 2015 Aug;282(15):2806-26. doi: 10.1111/febs.13305. Epub 2015 May 9. FEBS J. 2015. PMID: 25899482 Free PMC article. Review.
References
-
- Farrer MJ. Genetics of Parkinson disease: paradigm shifts and future prospects. Nat Rev Genet. 2006;7:306–318. - PubMed
-
- Marin I. The Parkinson Disease Gene LRRK2: Evolutionary and Structural Insights. Mol Biol Evol. 2006;23:2423–2433. - PubMed
-
- Mata IF, Wedemeyer WJ, Farrer MJ, Taylor JP, Gallo KA. LRRK2 in Parkinson's disease: protein domains and functional insights. Trends Neurosci. 2006;29:286–293. - PubMed
-
- Gloeckner CJ, Kinkl N, Schumacher A, Braun RJ, O'Neill E, Meitinger T, et al. The Parkinson disease causing LRRK2 mutation I2020T is associated with increased kinase activity. Hum Mol Genet. 2006;15:223–232. - PubMed
-
- Greggio E, Jain S, Kingsbury A, Bandopadhyay R, Lewis P, Kaganovich A, et al. Kinase activity is required for the toxic effects of mutant LRRK2/dardarin. Neurobiol Dis. 2006;23:329–341. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases