Molecular characterization of TMPRSS2-ERG gene fusion in the NCI-H660 prostate cancer cell line: a new perspective for an old model
- PMID: 17401460
- PMCID: PMC1838578
- DOI: 10.1593/neo.07103
Molecular characterization of TMPRSS2-ERG gene fusion in the NCI-H660 prostate cancer cell line: a new perspective for an old model
Abstract
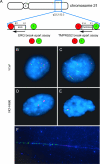
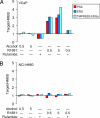
Recent studies have established that a significant fraction of prostate cancers harbor a signature gene fusion between the 5' region of androgen-regulated TMPRSS2 and an ETS family transcription factor, most commonly ERG. Studies on the molecular mechanisms and functional consequences of this important chromosomal rearrangement are currently limited to the VCaP cell line derived from a vertebral bone metastasis of a hormone-refractory prostate tumor. Here we report on the NCI-H660 cell line, derived from a metastatic site of an extrapulmonary small cell carcinoma arising from the prostate. NCI-H660 harbors TMPRSS2-ERG fusion with a homozygous intronic deletion between TMPRSS2 and ERG. We demonstrate this by real-time quantitative polymerase chain reaction, a two-stage dual-color interphase fluorescence in situ hybridization (FISH) assay testing for TMPRSS2 and ERG break-aparts, and single-nucleotide polymorphism oligonucleotide arrays. The deletion is consistent with the common intronic deletion found on chromosome 21q22.2-3 in human prostate cancer samples. We demonstrate the physical juxtaposition of TMPRSS2 and ERG on the DNA level by fiber FISH. The androgen receptor-negative NCI-H660 cell line expresses ERG in an androgen-independent fashion. This in vitro model system has the potential to provide important pathobiologic insights into TMPRSS2-ERG fusion prostate cancer.
Figures
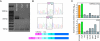

Similar articles
-
Morphological features of TMPRSS2-ERG gene fusion prostate cancer.J Pathol. 2007 May;212(1):91-101. doi: 10.1002/path.2154. J Pathol. 2007. PMID: 17385188
-
TMPRSS2:ERG fusion by translocation or interstitial deletion is highly relevant in androgen-dependent prostate cancer, but is bypassed in late-stage androgen receptor-negative prostate cancer.Cancer Res. 2006 Nov 15;66(22):10658-63. doi: 10.1158/0008-5472.CAN-06-1871. Cancer Res. 2006. PMID: 17108102
-
Noninvasive detection of TMPRSS2:ERG fusion transcripts in the urine of men with prostate cancer.Neoplasia. 2006 Oct;8(10):885-8. doi: 10.1593/neo.06625. Neoplasia. 2006. PMID: 17059688 Free PMC article.
-
[The progress of TMPRSS2-ETS gene fusions and their mechanism in prostate cancer].Yi Chuan. 2011 Feb;33(2):117-22. doi: 10.3724/sp.j.1005.2011.00117. Yi Chuan. 2011. PMID: 21377967 Review. Chinese.
-
Prevalence and clinical application of TMPRSS2-ERG fusion in Asian prostate cancer patients: a large-sample study in Chinese people and a systematic review.Asian J Androl. 2020 Mar-Apr;22(2):200-207. doi: 10.4103/aja.aja_45_19. Asian J Androl. 2020. PMID: 31210145 Free PMC article.
Cited by
-
Detection of TMPRSS2-ERG fusion gene in benign prostatic hyperplasia.Tumour Biol. 2014 Oct;35(10):9597-602. doi: 10.1007/s13277-014-2250-0. Epub 2014 Jun 25. Tumour Biol. 2014. PMID: 24961351
-
Increased nuclear factor I/B expression in prostate cancer correlates with AR expression.Prostate. 2020 Sep;80(13):1058-1070. doi: 10.1002/pros.24019. Epub 2020 Jul 21. Prostate. 2020. PMID: 32692871 Free PMC article.
-
ERG gene rearrangements are common in prostatic small cell carcinomas.Mod Pathol. 2011 Jun;24(6):820-8. doi: 10.1038/modpathol.2011.7. Epub 2011 Feb 18. Mod Pathol. 2011. PMID: 21336263 Free PMC article.
-
Golgi protein GOLM1 is a tissue and urine biomarker of prostate cancer.Neoplasia. 2008 Nov;10(11):1285-94. doi: 10.1593/neo.08922. Neoplasia. 2008. PMID: 18953438 Free PMC article.
-
Organoid cultures derived from patients with advanced prostate cancer.Cell. 2014 Sep 25;159(1):176-187. doi: 10.1016/j.cell.2014.08.016. Epub 2014 Sep 4. Cell. 2014. PMID: 25201530 Free PMC article.
References
-
- Tomlins SA, Rhodes DR, Perner S, Dhanasekaran SM, Mehra R, Sun XW, Varambally S, Cao X, Tchinda J, Kuefer R, et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science. 2005;310:644–648. - PubMed
-
- Tomlins SA, Mehra R, Rhodes DR, Smith LR, Roulston D, Helgeson BE, Cao X, Wei JT, Rubin MA, Shah RB, et al. TMPRSS2:ETV4 gene fusions define a third molecular subtype of prostate cancer. Cancer Res. 2006;66:3396–3400. - PubMed
-
- Perner S, Demichelis F, Beroukhim R, Schmidt FH, Mosquera JM, Setlur S, Tchinda J, Tomlins SA, Hofer MD, Pienta KG, et al. TMPRSS2:ERG fusion-associated deletions provide insight into the heterogeneity of prostate cancer. Cancer Res. 2006;66:8337–8341. - PubMed
-
- Demichelis F, Fall K, Perner S, Andren O, Schmidt F, Setlur SR, Hoshida Y, Mosquera J-M, Pawitan Y, Lee C, et al. TMPRSS2:ERG gene fusion associated with lethal prostate cancer. Oncogene. 2007 2007 Jan 22 [Epub ahead of print] - PubMed
-
- Korenchuk S, Lehr JE, MClean L, Lee YG, Whitney S, Vessella R, Lin DL, Pienta KJ. VCaP, a cell-based model system of human prostate cancer. In Vivo. 2001;15:163–168. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
