Discovery of tMAC: a Drosophila testis-specific meiotic arrest complex paralogous to Myb-Muv B
- PMID: 17403774
- PMCID: PMC1847709
- DOI: 10.1101/gad.1516607
Discovery of tMAC: a Drosophila testis-specific meiotic arrest complex paralogous to Myb-Muv B
Abstract
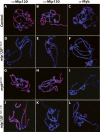
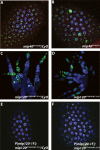
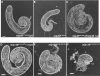
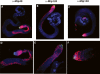
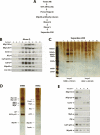
The Drosophila Myb-Muv B (MMB)/dREAM complex regulates gene expression and DNA replication site-specifically, but its activities in vivo have not been thoroughly explored. In ovarian amplification-stage follicle cell nuclei, the largest subunit, Mip130, is a negative regulator of replication, whereas another subunit, Myb, is a positive regulator. Here, we identified a mutation in mip40 and generated a mutation in mip120, two additional MMB subunits. Both mutants were viable, but mip120 mutants had many complex phenotypes including shortened longevity and severe eye defects. mip40 mutant females had severely reduced fertility, whereas mip120 mutant females were sterile, substantiating ovarian regulatory role(s) for MMB. Myb accumulation and binding to polytene chromosomes was dependent on the core factors of the MMB complex. In contrast to the documented mip130 mutant phenotypes, both mip40 and mip120 mutant males were sterile. We purified Mip40-containing complexes from testis nuclear extracts and identified tMAC, a new testis-specific meiotic arrest complex that contained Mip40, Caf1/p55, the Mip130 family member, Always early (Aly), and a Mip120 family member, Tombola (Tomb). Together, these data demonstrate that MMB serves diverse roles in different developmental pathways, and members of MMB can be found in alternative, noninteracting complexes in different cell types.
Figures

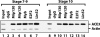

References
-
- Ashburner M. Drosophila: A laboratory manual. Cold Spring Harbor Laboratory Press; Cold Spring Harbor, NY: 1989.
-
- Beall E.L., Manak J.R., Zhou S., Bell M., Lipsick J.S., Botchan M.R., Manak J.R., Zhou S., Bell M., Lipsick J.S., Botchan M.R., Zhou S., Bell M., Lipsick J.S., Botchan M.R., Bell M., Lipsick J.S., Botchan M.R., Lipsick J.S., Botchan M.R., Botchan M.R. Role for a Drosophila Myb-containing protein complex in site-specific DNA replication. Nature. 2002;420:833–837. - PubMed
-
- Boxem M., van den Heuvel S., van den Heuvel S. C. elegans class B synthetic multivulva genes act in G(1) regulation. Curr. Biol. 2002;12:906–911. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
