In silico pathway reconstruction: Iron-sulfur cluster biogenesis in Saccharomyces cerevisiae
- PMID: 17408500
- PMCID: PMC1839888
- DOI: 10.1186/1752-0509-1-10
In silico pathway reconstruction: Iron-sulfur cluster biogenesis in Saccharomyces cerevisiae
Abstract
Background: Current advances in genomics, proteomics and other areas of molecular biology make the identification and reconstruction of novel pathways an emerging area of great interest. One such class of pathways is involved in the biogenesis of Iron-Sulfur Clusters (ISC).
Results: Our goal is the development of a new approach based on the use and combination of mathematical, theoretical and computational methods to identify the topology of a target network. In this approach, mathematical models play a central role for the evaluation of the alternative network structures that arise from literature data-mining, phylogenetic profiling, structural methods, and human curation. As a test case, we reconstruct the topology of the reaction and regulatory network for the mitochondrial ISC biogenesis pathway in S. cerevisiae. Predictions regarding how proteins act in ISC biogenesis are validated by comparison with published experimental results. For example, the predicted role of Arh1 and Yah1 and some of the interactions we predict for Grx5 both matches experimental evidence. A putative role for frataxin in directly regulating mitochondrial iron import is discarded from our analysis, which agrees with also published experimental results. Additionally, we propose a number of experiments for testing other predictions and further improve the identification of the network structure.
Conclusion: We propose and apply an iterative in silico procedure for predictive reconstruction of the network topology of metabolic pathways. The procedure combines structural bioinformatics tools and mathematical modeling techniques that allow the reconstruction of biochemical networks. Using the Iron Sulfur cluster biogenesis in S. cerevisiae as a test case we indicate how this procedure can be used to analyze and validate the network model against experimental results. Critical evaluation of the obtained results through this procedure allows devising new wet lab experiments to confirm its predictions or provide alternative explanations for further improving the models.
Figures

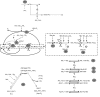
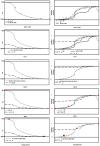
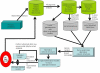
Similar articles
-
How Escherichia coli and Saccharomyces cerevisiae build Fe/S proteins.Adv Microb Physiol. 2005;50:41-101. doi: 10.1016/S0065-2911(05)50002-X. Adv Microb Physiol. 2005. PMID: 16221578
-
Predictive reconstruction of the mitochondrial iron-sulfur cluster assembly metabolism. II. Role of glutaredoxin Grx5.Proteins. 2004 Nov 15;57(3):481-92. doi: 10.1002/prot.20228. Proteins. 2004. PMID: 15382238
-
A dynamic model of the proteins that form the initial iron-sulfur cluster biogenesis machinery in yeast mitochondria.Protein J. 2013 Mar;32(3):183-96. doi: 10.1007/s10930-013-9475-4. Protein J. 2013. PMID: 23463383
-
The role of mitochondria in cellular iron-sulfur protein biogenesis and iron metabolism.Biochim Biophys Acta. 2012 Sep;1823(9):1491-508. doi: 10.1016/j.bbamcr.2012.05.009. Epub 2012 May 15. Biochim Biophys Acta. 2012. PMID: 22609301 Review.
-
Mitochondrial iron-sulfur protein biogenesis and human disease.Biochimie. 2014 May;100:61-77. doi: 10.1016/j.biochi.2014.01.010. Epub 2014 Jan 23. Biochimie. 2014. PMID: 24462711 Review.
Cited by
-
Biblio-MetReS: a bibliometric network reconstruction application and server.BMC Bioinformatics. 2011 Oct 5;12:387. doi: 10.1186/1471-2105-12-387. BMC Bioinformatics. 2011. PMID: 21975133 Free PMC article.
-
A mathematical model for strigolactone biosynthesis in plants.Front Plant Sci. 2022 Sep 2;13:979162. doi: 10.3389/fpls.2022.979162. eCollection 2022. Front Plant Sci. 2022. PMID: 36119618 Free PMC article.
-
A kinetic model of iron trafficking in growing Saccharomyces cerevisiae cells; applying mathematical methods to minimize the problem of sparse data and generate viable autoregulatory mechanisms.PLoS Comput Biol. 2023 Dec 19;19(12):e1011701. doi: 10.1371/journal.pcbi.1011701. eCollection 2023 Dec. PLoS Comput Biol. 2023. PMID: 38113197 Free PMC article.
-
Metabolic pathway reconstruction of eugenol to vanillin bioconversion in Aspergillus niger.Bioinformation. 2010 Jan 23;4(7):320-5. doi: 10.6026/97320630004320. Bioinformation. 2010. PMID: 20978605 Free PMC article.
-
A comprehensive mechanistic model of iron metabolism in Saccharomyces cerevisiae.Metallomics. 2019 Nov 1;11(11):1779-1799. doi: 10.1039/c9mt00199a. Epub 2019 Sep 18. Metallomics. 2019. PMID: 31531508 Free PMC article.
References
-
- Alves R, Antunes F, Salvador A. Tools for kinetic modeling of biochemical networks. Nat Biotechnol. 2006;24:667–672. - PubMed
-
- Francke C, Siezen RJ, Teusink B. Reconstructing the metabolic network of a bacterium from its genome. Trends Microbiol. 2005;13:550–558. - PubMed
-
- Ideker T. A systems approach to discovering signaling and regulatory pathways – or, how to digest large interaction networks into relevant pieces. Adv Exp Med Biol. 2004;547:21–30. - PubMed
-
- Ideker T, Winslow LR, Lauffenburger AD. Bioengineering and systems biology. Ann Biomed Eng. 2006;34:257–264. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

