LIN-61, one of two Caenorhabditis elegans malignant-brain-tumor-repeat-containing proteins, acts with the DRM and NuRD-like protein complexes in vulval development but not in certain other biological processes
- PMID: 17409073
- PMCID: PMC1893064
- DOI: 10.1534/genetics.106.069633
LIN-61, one of two Caenorhabditis elegans malignant-brain-tumor-repeat-containing proteins, acts with the DRM and NuRD-like protein complexes in vulval development but not in certain other biological processes
Abstract
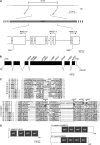
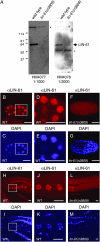
Vulval development in Caenorhabiditis elegans is inhibited by the redundant functions of the synthetic multivulva (synMuv) genes. At least 26 synMuv genes have been identified, many of which appear to act via transcriptional repression. Here we report the molecular identification of the class B synMuv gene lin-61, which encodes a protein composed of four malignant brain tumor (MBT) repeats. MBT repeats, domains of approximately 100 amino acids, have been found in multiple copies in a number of transcriptional repressors, including Polycomb-group proteins. MBT repeats are important for the transcriptional repression mediated by these proteins and in some cases have been shown to bind modified histones. C. elegans contains one other MBT-repeat-containing protein, MBTR-1. We demonstrate that a deletion allele of mbtr-1 does not cause a synMuv phenotype nor does mbtr-1 appear to act redundantly with or in opposition to lin-61. We further show that lin-61 is phenotypically and biochemically distinct from other class B synMuv genes. Our data indicate that while the class B synMuv genes act together to regulate vulval development, lin-61 functions separately from some class B synMuv proteins in other biological processes.
Figures




References
-
- Andersen, E. C., X. Lu and H. R. Horvitz, 2006. C. elegans ISWI and NURF301 antagonize an Rb-like pathway in the determination of multiple cell fates. Development 133: 2695–2704. - PubMed
-
- Bannister, A. J., P. Zegerman, J. F. Partridge, E. A. Miska, J. O. Thomas et al., 2001. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410: 120–124. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

