Enumeration of oligomerization states of membrane proteins in living cells by homo-FRET spectroscopy and microscopy: theory and application
- PMID: 17416632
- PMCID: PMC1852368
- DOI: 10.1529/biophysj.106.099424
Enumeration of oligomerization states of membrane proteins in living cells by homo-FRET spectroscopy and microscopy: theory and application
Abstract
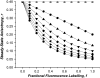
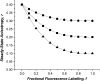
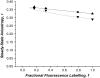
Protein-protein interactions play a pivotal role in biological signaling networks. It is highly desirable to perform experiments that can directly assess the oligomerization state and degree of oligomerization of biological macromolecules in their native environment. Homo-FRET depends on the inverse sixth power of separation between interacting like fluorophores on the nanometer scale and is therefore sensitive to protein oligomerization. Homo-FRET is normally detected by steady-state or time-resolved fluorescence anisotropy measurements. Here we show by theory and simulation that an examination of the extent of homotransfer as measured by steady-state fluorescence anisotropy as a function of fluorophore labeling (or photodepletion) gives valuable information on the oligomerization state of self-associating proteins. We examine random distributions of monomers, dilute solutions of oligomers, and concentrated solutions of oligomers. The theory is applied to literature data on band 3 protein dimers in membranes, GPI-linked protein trimers in "rafts," and clustered GFP-tagged epidermal growth factor receptors in cell membranes to illustrate the general utility and applicability of our analytical approach.
Figures




Similar articles
-
Exploring homo-FRET to quantify the oligomer stoichiometry of membrane-bound proteins involved in a cooperative partition equilibrium.Phys Chem Chem Phys. 2014 Sep 14;16(34):18105-17. doi: 10.1039/c4cp00060a. Phys Chem Chem Phys. 2014. PMID: 24722583
-
Analysis of EGF receptor oligomerization by homo-FRET.Methods Cell Biol. 2013;117:305-21. doi: 10.1016/B978-0-12-408143-7.00016-5. Methods Cell Biol. 2013. PMID: 24143984
-
Red-edge anisotropy microscopy enables dynamic imaging of homo-FRET between green fluorescent proteins in cells.J Struct Biol. 2004 Jul;147(1):62-9. doi: 10.1016/j.jsb.2003.10.013. J Struct Biol. 2004. PMID: 15109606
-
Förster resonance energy transfer microscopy and spectroscopy for localizing protein-protein interactions in living cells.Cytometry A. 2013 Sep;83(9):780-93. doi: 10.1002/cyto.a.22321. Epub 2013 Jun 27. Cytometry A. 2013. PMID: 23813736 Free PMC article. Review.
-
Fluorescence anisotropy imaging microscopy for homo-FRET in living cells.Methods Cell Biol. 2008;85:395-414. doi: 10.1016/S0091-679X(08)85017-0. Methods Cell Biol. 2008. PMID: 18155472 Review.
Cited by
-
Fibroblast growth factor receptor 5 (FGFR5) is a co-receptor for FGFR1 that is up-regulated in beta-cells by cytokine-induced inflammation.J Biol Chem. 2018 Nov 2;293(44):17218-17228. doi: 10.1074/jbc.RA118.003036. Epub 2018 Sep 14. J Biol Chem. 2018. PMID: 30217817 Free PMC article.
-
Distribution of resting and ligand-bound ErbB1 and ErbB2 receptor tyrosine kinases in living cells using number and brightness analysis.Proc Natl Acad Sci U S A. 2010 Sep 21;107(38):16524-9. doi: 10.1073/pnas.1002642107. Epub 2010 Sep 2. Proc Natl Acad Sci U S A. 2010. PMID: 20813958 Free PMC article.
-
Fluorescence-based approaches for monitoring membrane receptor oligomerization.J Biosci. 2018 Jul;43(3):463-469. J Biosci. 2018. PMID: 30002266 Review.
-
Amphiphilic Gold Nanoparticles: A Biomimetic Tool to Gain Mechanistic Insights into Peptide-Lipid Interactions.Membranes (Basel). 2022 Jun 29;12(7):673. doi: 10.3390/membranes12070673. Membranes (Basel). 2022. PMID: 35877876 Free PMC article. Review.
-
When one plus one does not equal two: fluorescence anisotropy in aggregates and multiply labeled proteins.Biophys J. 2014 Apr 1;106(7):1457-66. doi: 10.1016/j.bpj.2014.02.020. Biophys J. 2014. PMID: 24703307 Free PMC article.
References
-
- Ullrich, A., and J. Schlessinger. 1990. Signal transduction by receptors with tyrosine kinase activity. Cell. 61:203–212. - PubMed
-
- Metzger, H. 1992. Transmembrane signaling: the joy of aggregation. J. Immunol. 149:1477–1487. - PubMed
-
- Yarden, Y., and J. Schlessinger. 1987. Self-phosphorylation of epidermal growth factor receptor: evidence for a model of intermolecular allosteric activation. Biochemistry. 26:1434–1442. - PubMed
-
- Jovin, T. M., and W. L. Vaz. 1989. Rotational and translational diffusion in membranes measured by fluorescence and phosphorescence methods. Methods Enzymol. 172:471–513. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

