CpG methylation is targeted to transcription units in an invertebrate genome
- PMID: 17420183
- PMCID: PMC1855171
- DOI: 10.1101/gr.6163007
CpG methylation is targeted to transcription units in an invertebrate genome
Abstract
DNA is methylated at the dinucleotide CpG in genomes of a wide range of plants and animals. Among animals, variable patterns of genomic CpG methylation have been described, ranging from undetectable levels (e.g., in Caenorhabditis elegans) to high levels of global methylation in the vertebrates. The most frequent pattern in invertebrate animals, however, is mosaic methylation, comprising domains of methylated DNA interspersed with unmethylated domains. To understand the origin of mosaic DNA methylation patterns, we examined the distribution of DNA methylation in the Ciona intestinalis genome. Bisulfite sequencing and computational analysis revealed methylated domains with sharp boundaries that strongly colocalize with approximately 60% of transcription units. By contrast, promoters, intergenic DNA, and transposons are not preferentially targeted by DNA methylation. Methylated transcription units include evolutionarily conserved genes, whereas the most highly expressed genes preferentially belong to the unmethylated fraction. The results lend support to the hypothesis that CpG methylation functions to suppress spurious transcriptional initiation within infrequently transcribed genes.
Figures
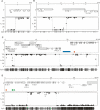

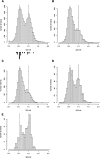

References
-
- Bird A.P. CpG-rich islands and the function of DNA methylation. Nature. 1986;321:209–213. - PubMed
-
- Bird A.P. CpG islands as gene markers in the vertebrate nucleus. Trends Genet. 1987;3:342–347.
-
- Bird A.P. Gene number, noise reduction and biological complexity. Trends Genet. 1995;11:94–100. - PubMed
-
- Bird A. DNA methylation patterns and epigenetic memory. Genes & Dev. 2002;16:6–21. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
