A primary assembly of a bovine haplotype block map based on a 15,036-single-nucleotide polymorphism panel genotyped in holstein-friesian cattle
- PMID: 17435229
- PMCID: PMC1894606
- DOI: 10.1534/genetics.106.069369
A primary assembly of a bovine haplotype block map based on a 15,036-single-nucleotide polymorphism panel genotyped in holstein-friesian cattle
Abstract
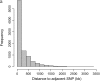
Analysis of data on 1000 Holstein-Friesian bulls genotyped for 15,036 single-nucleotide polymorphisms (SNPs) has enabled genomewide identification of haplotype blocks and tag SNPs. A final subset of 9195 SNPs in Hardy-Weinberg equilibrium and mapped on autosomes on the bovine sequence assembly (release Btau 3.1) was used in this study. The average intermarker spacing was 251.8 kb. The average minor allele frequency (MAF) was 0.29 (0.05-0.5). Following recent precedents in human HapMap studies, a haplotype block was defined where 95% of combinations of SNPs within a region are in very high linkage disequilibrium. A total of 727 haplotype blocks consisting of > or =3 SNPs were identified. The average block length was 69.7 +/- 7.7 kb, which is approximately 5-10 times larger than in humans. These blocks comprised a total of 2964 SNPs and covered 50,638 kb of the sequence map, which constitutes 2.18% of the length of all autosomes. A set of tag SNPs, which will be useful for further fine-mapping studies, has been identified. Overall, the results suggest that as many as 75,000-100,000 tag SNPs would be needed to track all important haplotype blocks in the bovine genome. This would require approximately 250,000 SNPs in the discovery phase.
Figures



Similar articles
-
High-resolution haplotype block structure in the cattle genome.BMC Genet. 2009 Apr 24;10:19. doi: 10.1186/1471-2156-10-19. BMC Genet. 2009. PMID: 19393054 Free PMC article.
-
[Comparison of minor allele frequencies and haplotype frequencies for single nucleotide polymorphisms in ROR2 gene using HapMap data for Chinese Hans in Beijing and Yoruban in Ibadan in Nigeria].Beijing Da Xue Xue Bao Yi Xue Ban. 2011 Dec 18;43(6):785-91. Beijing Da Xue Xue Bao Yi Xue Ban. 2011. PMID: 22178821 Chinese.
-
The pattern of linkage disequilibrium in German Holstein cattle.Anim Genet. 2010 Aug;41(4):346-56. doi: 10.1111/j.1365-2052.2009.02011.x. Epub 2010 Jan 3. Anim Genet. 2010. PMID: 20055813
-
Tag SNP selection for association studies.Genet Epidemiol. 2004 Dec;27(4):365-74. doi: 10.1002/gepi.20028. Genet Epidemiol. 2004. PMID: 15372618 Review.
-
[Analysis and application of SNP and haplotype in the human genome].Yi Chuan Xue Bao. 2005 Aug;32(8):879-89. Yi Chuan Xue Bao. 2005. PMID: 16231744 Review. Chinese.
Cited by
-
Mapping of a milk production quantitative trait locus to a 1.056 Mb region on bovine chromosome 5 in the Fleckvieh dual purpose cattle breed.Genet Sel Evol. 2011 Feb 24;43(1):8. doi: 10.1186/1297-9686-43-8. Genet Sel Evol. 2011. PMID: 21349166 Free PMC article.
-
Gene mapping in the wild with SNPs: guidelines and future directions.Genetica. 2009 May;136(1):97-107. doi: 10.1007/s10709-008-9317-z. Epub 2008 Sep 9. Genetica. 2009. PMID: 18780148
-
High-resolution haplotype block structure in the cattle genome.BMC Genet. 2009 Apr 24;10:19. doi: 10.1186/1471-2156-10-19. BMC Genet. 2009. PMID: 19393054 Free PMC article.
-
Discovery of novel genetic networks associated with 19 economically important traits in beef cattle.Int J Biol Sci. 2009 Jul 29;5(6):528-42. doi: 10.7150/ijbs.5.528. Int J Biol Sci. 2009. PMID: 19727437 Free PMC article.
-
Genetic and haplotypic structure in 14 European and African cattle breeds.Genetics. 2007 Oct;177(2):1059-70. doi: 10.1534/genetics.107.075804. Epub 2007 Aug 24. Genetics. 2007. PMID: 17720924 Free PMC article.
References
-
- Barrett, J. C., B. Fry, J. Maller and M. J. Daly, 2005. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21 263–265. - PubMed
-
- Cardon, L. R., and G. R. Abecasis, 2003. Using haplotype blocks to map human complex trait loci. Trends Genet. 19 135–140. - PubMed
-
- Cohen, M., M. Reichenstein, A. Everts-van der Wind, J. Heon-Lee, M. Shani et al., 2004. Cloning and characterization of FAM13A1—a gene near a milk protein QTL on BTA6: evidence for population-wide linkage disequilibrium in Israeli Holsteins. Genomics 84 374–383. - PubMed
-
- Craig, D. W., and D. A. Stephan, 2005. Applications of whole-genome high-density SNP genotyping. Expert Rev. Mol. Diagn. 5 159–170. - PubMed
-
- Daly, M. J., J. D. Rioux, S. F. Schaffner, T. J. Hudson and E. S. Lander, 2001. High-resolution haplotype structure in the human genome. Nat. Genet. 29 229–232. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

