Multiple trans-sensing interactions affect meiotically heritable epigenetic states at the maize pl1 locus
- PMID: 17435245
- PMCID: PMC1894611
- DOI: 10.1534/genetics.107.072496
Multiple trans-sensing interactions affect meiotically heritable epigenetic states at the maize pl1 locus
Abstract
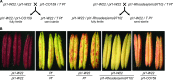
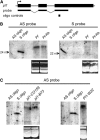
Interactions between specific maize purple plant1 (pl1) alleles result in heritable changes of gene regulation that are manifested as differences in anthocyanin pigmentation. Transcriptionally repressed states of Pl1-Rhoades alleles (termed Pl') are remarkably stable and invariably facilitate heritable changes of highly expressed states (termed Pl-Rh) in Pl'/Pl-Rh plants. However, Pl' can revert to Pl-Rh when hemizygous, when heterozygous with pl1 alleles other than Pl1-Rhoades, or in the absence of trans-acting factors required to maintain repressed states. Cis-linked features of Pl1-Rhoades responsible for these trans-sensing behaviors remain unknown. Here, genetic tests of a pl1 allelic series identify two potentially separate cis-linked features: one facilitating repression of Pl-Rh and another stabilizing Pl' in trans. Neither function is affected in ethyl-methanesulfonate-induced Pl1-Rhoades derivatives that produce truncated PL1 peptides, indicating that PL1 is unlikely to mediate trans interactions. Both functions, however, are impaired in a spontaneous Pl1-Rhoades derivative that fails to produce detectable pl1 RNA. Pl'-like states can also repress expression of a pl1-W22 allele, but this repression is not meiotically heritable. As the Pl' state is not associated with unique small RNA species representing the pl1-coding region, the available data suggest that interactions between elements required for transcription underlie Pl1-Rhoades epigenetic behaviors.
Figures



References
-
- Alleman, M., L. Sidorenko, K. McGinnis, V. Seshadri, J. E. Dorweiler et al., 2006. An RNA-dependent RNA polymerase is required for paramutation in maize. Nature 442 295–298. - PubMed
-
- Brink, R. A., 1964. Genetic repression of R action in maize, pp. 183–230 in The Role of Chromosomes in Development, edited by M. Locke. Academic Press, New York.
-
- Brodersen, P., and O. Voinnet, 2006. The diversity of RNA silencing pathways in plants. Trends Genet. 22 268–280. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

