Elution of tightly bound solutes from concanavalin A Sepharose. Factors affecting the desorption of cottonmouth venom glycoproteins
- PMID: 17449042
- PMCID: PMC2040237
- DOI: 10.1016/j.chroma.2007.03.126
Elution of tightly bound solutes from concanavalin A Sepharose. Factors affecting the desorption of cottonmouth venom glycoproteins
Abstract
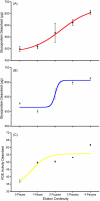
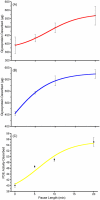
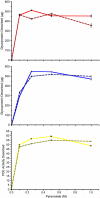
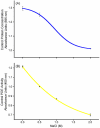
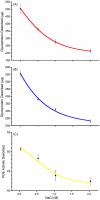
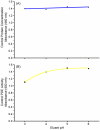
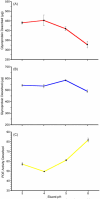
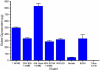
Some glycoproteins bind so tightly to concanavalin A Sepharose that common desorption techniques are ineffective, so a systematic exploration of factors affecting desorption of cottonmouth venom glycoproteins was undertaken. Glycoprotein desorption is greatly improved by introducing up to four pauses of 5-10 min duration into the elution step. Eluent concentrations above 250 mM methylglucoside or 500 mM methyl-mannoside reduced glycoprotein desorption. Eluent NaCl diminished glycoprotein desorption. Most venom glycoproteins desorb more readily as pH diminishes from 6.0 to 4.0, but phosphodiesterase shows the opposite pattern. Eluents recommended by the supplier for desorbing solutes or for column cleaning were ineffectual.
Figures

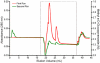
Similar articles
-
Improved concanavalin A-Sepharose elution by specific readsorption of glycoproteins.J Chromatogr. 1983 May 20;261(1):77-82. doi: 10.1016/s0021-9673(01)87920-6. J Chromatogr. 1983. PMID: 6575973
-
Venom exonuclease (phosphodiesterase) immobilized on concanavalin-A-sepharose.Biochem Biophys Res Commun. 1974 Mar 25;57(2):463-8. doi: 10.1016/0006-291x(74)90954-1. Biochem Biophys Res Commun. 1974. PMID: 4364241 No abstract available.
-
Phospholipase A2 in the venom of three cottonmouth snakes.Toxicon. 2017 Sep 1;135:84-92. doi: 10.1016/j.toxicon.2017.06.010. Epub 2017 Jun 17. Toxicon. 2017. PMID: 28633930
-
Purification of the glycoprotein allergen Ag7 from mugwort pollen by concanavalin A affinity chromatography.J Biotechnol. 1990 Nov;16(3-4):305-16. doi: 10.1016/0168-1656(90)90044-c. J Biotechnol. 1990. PMID: 1366940
-
Improved performance of column chromatography by arginine: dye-affinity chromatography.Protein Expr Purif. 2007 Apr;52(2):410-4. doi: 10.1016/j.pep.2006.10.005. Epub 2006 Oct 25. Protein Expr Purif. 2007. PMID: 17126030
Cited by
-
Characterization of the human submandibular/sublingual saliva glycoproteome using lectin affinity chromatography coupled to multidimensional protein identification technology.J Proteome Res. 2011 Nov 4;10(11):5031-46. doi: 10.1021/pr200505t. Epub 2011 Oct 13. J Proteome Res. 2011. PMID: 21936497 Free PMC article.
-
The Novel Type 1 Fimbriae FimH Receptor Calreticulin Plays a Role in Salmonella Host Specificity.Front Cell Infect Microbiol. 2017 Jul 19;7:326. doi: 10.3389/fcimb.2017.00326. eCollection 2017. Front Cell Infect Microbiol. 2017. PMID: 28770174 Free PMC article.
-
Carboxyl-terminal truncations alter the activity of the human α-galactosidase A.PLoS One. 2015 Feb 26;10(2):e0118341. doi: 10.1371/journal.pone.0118341. eCollection 2015. PLoS One. 2015. PMID: 25719393 Free PMC article.
References
-
- Fountoulakis M, Juranville JF. J. Biochem. Biophys. Methods. 1993;27:127. - PubMed
-
- Hayman MJ, Crumpton MJ. Biochem. Biophys. Res. Commun. 1972;47:923. - PubMed
-
- Bishayee S, Farooqui AA, Bachhawat BK. Indian J. Biochem. Biophys. 1973;10:1. - PubMed
-
- Davey MW, Huang JW, Sulkowski E, Carter WA. J. Biol. Chem. 1974;249:6354. - PubMed
-
- Anderson JC. Biochim. Biophys. Acta. 1975;379:444. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

