NMR studies in dodecylphosphocholine of a fragment containing the seventh transmembrane helix of a G-protein-coupled receptor from Saccharomyces cerevisiae
- PMID: 17449670
- PMCID: PMC1896227
- DOI: 10.1529/biophysj.106.103770
NMR studies in dodecylphosphocholine of a fragment containing the seventh transmembrane helix of a G-protein-coupled receptor from Saccharomyces cerevisiae
Abstract
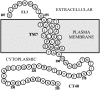
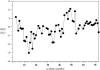
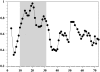
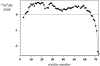
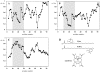
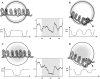
The structure and dynamics of a large segment of Ste2p, the G-protein-coupled alpha-factor receptor from yeast, were studied in dodecylphosphocholine (DPC) micelles using solution NMR spectroscopy. We investigated the 73-residue peptide EL3-TM7-CT40 consisting of the third extracellular loop 3 (EL3), the seventh transmembrane helix (TM7), and 40 residues from the cytosolic C-terminal domain (CT40). The structure reveals the presence of an alpha-helix in the segment encompassing residues 10-30, which is perturbed around the internal Pro-24 residue. Root mean-square deviation values of individually superimposed helical segments 10-20 and 25-30 were 0.91 +/- 0.33 A and 0.76 +/- 0.37 A, respectively. 15N-relaxation and residual dipolar coupling data support a rather stable fold for the TM7 part of EL3-TM7-CT40, whereas the EL3 and CT40 segments are more flexible. Spin-label data indicate that the TM7 helix integrates into DPC micelles but is flexible around the internal Pro-24 site, exposing residues 22-26 to solution and reveal a second site of interaction with the micelle within a region comprising residues 43-58, which forms part of a less well-defined nascent helix. These findings are discussed in light of previous studies in organic-aqueous solvent systems.
Figures





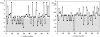
Similar articles
-
Structure of a double transmembrane fragment of a G-protein-coupled receptor in micelles.Biophys J. 2009 Apr 22;96(8):3187-96. doi: 10.1016/j.bpj.2009.01.012. Biophys J. 2009. PMID: 19383463 Free PMC article.
-
Expression and biophysical analysis of two double-transmembrane domain-containing fragments from a yeast G protein-coupled receptor.Biopolymers. 2008;90(2):117-30. doi: 10.1002/bip.20950. Biopolymers. 2008. PMID: 18260136
-
Structural characterization of triple transmembrane domain containing fragments of a yeast G protein-coupled receptor in an organic : aqueous environment by solution-state NMR spectroscopy.J Pept Sci. 2015 Mar;21(3):212-22. doi: 10.1002/psc.2750. Epub 2015 Feb 2. J Pept Sci. 2015. PMID: 25645975 Free PMC article.
-
Biosynthesis and NMR analysis of a 73-residue domain of a Saccharomyces cerevisiae G protein-coupled receptor.Biochemistry. 2005 Sep 6;44(35):11795-810. doi: 10.1021/bi0507231. Biochemistry. 2005. PMID: 16128581
-
Comparison of Experimental Approaches Used to Determine the Structure and Function of the Class D G Protein-Coupled Yeast α-Factor Receptor.Biomolecules. 2022 May 30;12(6):761. doi: 10.3390/biom12060761. Biomolecules. 2022. PMID: 35740886 Free PMC article. Review.
Cited by
-
Comparison of fragments comprising the first two helices of the human Y4 and the yeast Ste2p G-protein-coupled receptors.Biophys J. 2012 Aug 22;103(4):817-26. doi: 10.1016/j.bpj.2012.07.012. Biophys J. 2012. PMID: 22947943 Free PMC article.
-
Understanding GPCR Recognition and Folding from NMR Studies of Fragments.RSC Adv. 2018;8(18):9858-9870. doi: 10.1039/C8RA01520A. Epub 2018 Mar 9. RSC Adv. 2018. PMID: 29732143 Free PMC article.
-
Large multiple transmembrane domain fragments of a G protein-coupled receptor: biosynthesis, purification, and biophysical studies.Biopolymers. 2012;98(5):485-500. doi: 10.1002/bip.22122. Biopolymers. 2012. PMID: 23203693 Free PMC article.
-
Structure of a double transmembrane fragment of a G-protein-coupled receptor in micelles.Biophys J. 2009 Apr 22;96(8):3187-96. doi: 10.1016/j.bpj.2009.01.012. Biophys J. 2009. PMID: 19383463 Free PMC article.
-
Structural and dynamic properties of juxta-membrane segments of caveolin-1 and caveolin-2 at the membrane interface.Eur Biophys J. 2010 Jan;39(2):307-25. doi: 10.1007/s00249-009-0548-4. Epub 2009 Oct 22. Eur Biophys J. 2010. PMID: 19847421
References
-
- Mombaerts, P. 1999. Seven-transmembrane proteins as odorant and chemosensory receptors. Science. 286:707–711. - PubMed
-
- Lundstrom, K. 2005. The future of G protein-coupled receptors as targets in drug discovery. IDrugs. 8:909–913. - PubMed
-
- Thompson, M. D., W. M. Burnham, and D. E. Cole. 2005. The G protein-coupled receptors: pharmacogenetics and disease. Crit. Rev. Clin. Lab. Sci. 42:311–392. - PubMed
-
- Palczewski, K., T. Kumasaka, T. Hori, C. A. Behnke, H. Motoshima, B. A. Fox, I. Le Trong, D. C. Teller, T. Okada, R. E. Stenkamp, M. Yamamoto, and M. Miyano. 2000. Crystal structure of rhodopsin: a G protein-coupled receptor. Science. 289:739–745. - PubMed
-
- Flower, D. R. 1999. Modelling G-protein-coupled receptors for drug design. Biochim. Biophys. Acta. 1422:207–234. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

