The C-terminal loop of the homing endonuclease I-CreI is essential for site recognition, DNA binding and cleavage
- PMID: 17452357
- PMCID: PMC1904291
- DOI: 10.1093/nar/gkm183
The C-terminal loop of the homing endonuclease I-CreI is essential for site recognition, DNA binding and cleavage
Abstract
Meganucleases are sequence-specific endonucleases with large cleavage sites that can be used to induce efficient homologous gene targeting in cultured cells and plants. These enzymes open novel perspectives for genome engineering in a wide range of fields, including gene therapy. A new crystal structure of the I-CreI dimer without DNA has allowed the comparison with the DNA-bound protein. The C-terminal loop displays a different conformation, which suggests its implication in DNA binding. A site-directed mutagenesis study in this region demonstrates that whereas the C-terminal helix is negligible for DNA binding, the final C-terminal loop is essential in DNA binding and cleavage. We have identified two regions that comprise the Ser138-Lys139 and Lys142-Thr143 pairs whose double mutation affect DNA binding in vitro and abolish cleavage in vivo. However, the mutation of only one residue in these sites allows DNA binding in vitro and cleavage in vivo. These findings demonstrate that the C-terminal loop of I-CreI endonuclease plays a fundamental role in its catalytic mechanism and suggest this novel site as a region to take into account for engineering new endonucleases with tailored specificity.
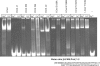
Figures




Similar articles
-
Engineered I-CreI derivatives cleaving sequences from the human XPC gene can induce highly efficient gene correction in mammalian cells.J Mol Biol. 2007 Aug 3;371(1):49-65. doi: 10.1016/j.jmb.2007.04.079. Epub 2007 May 10. J Mol Biol. 2007. PMID: 17561112
-
Engineering of large numbers of highly specific homing endonucleases that induce recombination on novel DNA targets.J Mol Biol. 2006 Jan 20;355(3):443-58. doi: 10.1016/j.jmb.2005.10.065. Epub 2005 Nov 15. J Mol Biol. 2006. PMID: 16310802
-
A combinatorial approach to create artificial homing endonucleases cleaving chosen sequences.Nucleic Acids Res. 2006;34(22):e149. doi: 10.1093/nar/gkl720. Epub 2006 Nov 27. Nucleic Acids Res. 2006. PMID: 17130168 Free PMC article.
-
Novel site-specific DNA endonucleases.Curr Opin Struct Biol. 1998 Feb;8(1):19-25. doi: 10.1016/s0959-440x(98)80005-5. Curr Opin Struct Biol. 1998. PMID: 9519292 Review.
-
The I-CreI meganuclease and its engineered derivatives: applications from cell modification to gene therapy.Protein Eng Des Sel. 2011 Jan;24(1-2):27-31. doi: 10.1093/protein/gzq083. Epub 2010 Nov 3. Protein Eng Des Sel. 2011. PMID: 21047873 Review.
Cited by
-
Molecular basis of engineered meganuclease targeting of the endogenous human RAG1 locus.Nucleic Acids Res. 2011 Jan;39(2):729-43. doi: 10.1093/nar/gkq801. Epub 2010 Sep 16. Nucleic Acids Res. 2011. PMID: 20846960 Free PMC article.
-
Improvements of nuclease and nickase gene modification techniques for the treatment of genetic diseases.Front Genome Ed. 2022 Jul 26;4:892769. doi: 10.3389/fgeed.2022.892769. eCollection 2022. Front Genome Ed. 2022. PMID: 35958050 Free PMC article. Review.
-
The Structural Basis of Asymmetry in DNA Binding and Cleavage as Exhibited by the I-SmaMI LAGLIDADG Meganuclease.J Mol Biol. 2016 Jan 16;428(1):206-220. doi: 10.1016/j.jmb.2015.12.005. Epub 2015 Dec 15. J Mol Biol. 2016. PMID: 26705195 Free PMC article.
-
Engineering a Nickase on the Homing Endonuclease I-DmoI Scaffold.J Biol Chem. 2015 Jul 24;290(30):18534-44. doi: 10.1074/jbc.M115.658666. Epub 2015 Jun 4. J Biol Chem. 2015. PMID: 26045557 Free PMC article.
-
Non-specific protein-DNA interactions control I-CreI target binding and cleavage.Nucleic Acids Res. 2012 Aug;40(14):6936-45. doi: 10.1093/nar/gks320. Epub 2012 Apr 11. Nucleic Acids Res. 2012. PMID: 22495931 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

