A complex system of small RNAs in the unicellular green alga Chlamydomonas reinhardtii
- PMID: 17470535
- PMCID: PMC1865491
- DOI: 10.1101/gad.1543507
A complex system of small RNAs in the unicellular green alga Chlamydomonas reinhardtii
Abstract
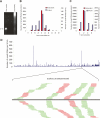
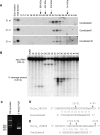
Endogenous small RNAs function in RNA interference (RNAi) pathways to control gene expression through mRNA cleavage, translational repression, or chromatin modification. Plants and animals contain many microRNAs (miRNAs) that play vital roles in development, including helping to specify cell type and tissue identity. To date, no miRNAs have been reported in unicellular organisms. Here we show that Chlamydomonas reinhardtii, a unicellular green alga, encodes many miRNAs. We also show that a Chlamydomonas miRNA can direct the cleavage of its target mRNA in vivo and in vitro. We further show that the expression of some miRNAs/Candidates increases or decreases during Chlamydomonas gametogenesis. In addition to miRNAs, Chlamydomonas harbors other types of small RNAs including phased small interfering RNAs (siRNAs) that are reminiscent of plant trans-acting siRNAs, as well as siRNAs originating from protein-coding genes and transposons. Our findings suggest that the miRNA pathway and some siRNA pathways are ancient mechanisms of gene regulation that evolved prior to the emergence of multicellularity.
Figures



Comment in
-
Expanding RNA physiology: microRNAs in a unicellular organism.Genes Dev. 2007 May 15;21(10):1153-6. doi: 10.1101/gad.1559707. Genes Dev. 2007. PMID: 17504934 Review. No abstract available.
References
-
- Allen E., Xie Z., Gustafson A.M., Sung G.H., Spatafora J.W., Carrington J.C., Xie Z., Gustafson A.M., Sung G.H., Spatafora J.W., Carrington J.C., Gustafson A.M., Sung G.H., Spatafora J.W., Carrington J.C., Sung G.H., Spatafora J.W., Carrington J.C., Spatafora J.W., Carrington J.C., Carrington J.C. Evolution of microRNA genes by inverted duplication of target gene sequences in Arabidopsis thaliana. Nat. Genet. 2004;36:1282–1290. - PubMed
-
- Allen E., Xie Z., Gustafson A.M., Carrington J.C., Xie Z., Gustafson A.M., Carrington J.C., Gustafson A.M., Carrington J.C., Carrington J.C. microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell. 2005;121:207–221. - PubMed
-
- Aravin A.A., Lagos-Quintana M., Yalcin A., Zavolan M., Marks D., Snyder B., Gaasterland T., Meyer J., Tuschl T., Lagos-Quintana M., Yalcin A., Zavolan M., Marks D., Snyder B., Gaasterland T., Meyer J., Tuschl T., Yalcin A., Zavolan M., Marks D., Snyder B., Gaasterland T., Meyer J., Tuschl T., Zavolan M., Marks D., Snyder B., Gaasterland T., Meyer J., Tuschl T., Marks D., Snyder B., Gaasterland T., Meyer J., Tuschl T., Snyder B., Gaasterland T., Meyer J., Tuschl T., Gaasterland T., Meyer J., Tuschl T., Meyer J., Tuschl T., Tuschl T. The small RNA profile during Drosophila melanogasterdevelopment. Dev. Cell. 2003;5:337–350. - PubMed
-
- Aravin A., Gaidatzis D., Pfeffer S., Lagos-Quintana M., Landgraf P., Iovino N., Morris P., Brownstein M.J., Kuramochi-Miyagawa S., Nakano T., Gaidatzis D., Pfeffer S., Lagos-Quintana M., Landgraf P., Iovino N., Morris P., Brownstein M.J., Kuramochi-Miyagawa S., Nakano T., Pfeffer S., Lagos-Quintana M., Landgraf P., Iovino N., Morris P., Brownstein M.J., Kuramochi-Miyagawa S., Nakano T., Lagos-Quintana M., Landgraf P., Iovino N., Morris P., Brownstein M.J., Kuramochi-Miyagawa S., Nakano T., Landgraf P., Iovino N., Morris P., Brownstein M.J., Kuramochi-Miyagawa S., Nakano T., Iovino N., Morris P., Brownstein M.J., Kuramochi-Miyagawa S., Nakano T., Morris P., Brownstein M.J., Kuramochi-Miyagawa S., Nakano T., Brownstein M.J., Kuramochi-Miyagawa S., Nakano T., Kuramochi-Miyagawa S., Nakano T., Nakano T., et al. A novel class of small RNAs bind to MILI protein in mouse testes. Nature. 2006;442:203–207. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
