Solution structure and backbone dynamics of the AF-6 PDZ domain/Bcr peptide complex
- PMID: 17473018
- PMCID: PMC2206671
- DOI: 10.1110/ps.062440607
Solution structure and backbone dynamics of the AF-6 PDZ domain/Bcr peptide complex
Abstract
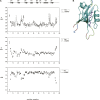
The human AF-6, a scaffold protein between cell membrane-associated proteins and the actin cytoskeleton, plays an important role in special cell-cell junctions and signal transduction. It can be phosphorylated by the protein kinase Bcr, which allows efficient binding of the C terminus of Bcr to the PDZ domain of AF-6 and consequently enhances the binding affinity of AF-6 to Ras. Formation of the AF-6, Bcr, and Ras ternary complex results in down-regulation of the Ras-mediated signal transduction pathway. To better understand the molecular basis for the recognition of the AF-6 PDZ domain and Bcr, we solve the solution structure of the AF-6 PDZ domain complexed with the C-terminal peptide of Bcr and explore the interactions between them in detail. Compared with previously reported structures, the complex exhibits a noncanonical binding mode of PDZ/peptide. Owing to the distinct residues involved in the AF-6 PDZ domain and Bcr peptide interaction, the interaction mode does not adapt to the existing classification rules that have been put forward, based on the ligand or the PDZ domain specificity. Furthermore, the PDZ domain of AF-6 can bind to the C terminus of Bcr efficiently after phosphorylation of AF-6 by the Bcr kinase. The phosphorylation may induce a conformational change of AF-6, which makes the binding surface on the PDZ domain accessible to Bcr for efficient binding. This study not only characterizes the structural details of the AF-6 PDZ/Bcr peptide complex, but also provides a potential target for future drug design and disease therapy.
Figures

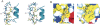
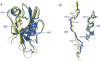
Similar articles
-
Interesting structural and dynamical behaviors exhibited by the AF-6 PDZ domain upon Bcr peptide binding.Biochemistry. 2007 Dec 25;46(51):15042-53. doi: 10.1021/bi701303p. Epub 2007 Dec 4. Biochemistry. 2007. PMID: 18052198
-
Solution structure of AF-6 PDZ domain and its interaction with the C-terminal peptides from Neurexin and Bcr.J Biol Chem. 2005 Apr 8;280(14):13841-7. doi: 10.1074/jbc.M411065200. Epub 2005 Jan 31. J Biol Chem. 2005. PMID: 15684424
-
The Bcr kinase downregulates Ras signaling by phosphorylating AF-6 and binding to its PDZ domain.Mol Cell Biol. 2003 Jul;23(13):4663-72. doi: 10.1128/MCB.23.13.4663-4672.2003. Mol Cell Biol. 2003. PMID: 12808105 Free PMC article.
-
Structures and target recognition modes of PDZ domains: recurring themes and emerging pictures.Biochem J. 2013 Oct 1;455(1):1-14. doi: 10.1042/BJ20130783. Biochem J. 2013. PMID: 24028161 Review.
-
AF-6/cno: neither a kinesin nor a myosin, but a bit of both.Trends Biochem Sci. 1995 Jul;20(7):265-6. doi: 10.1016/s0968-0004(00)89040-4. Trends Biochem Sci. 1995. PMID: 7667878 Review. No abstract available.
Cited by
-
Crystal structure of afadin PDZ domain-nectin-3 complex shows the structural plasticity of the ligand-binding site.Protein Sci. 2015 Mar;24(3):376-85. doi: 10.1002/pro.2628. Epub 2015 Jan 13. Protein Sci. 2015. PMID: 25534554 Free PMC article.
-
Interactions of Severe Acute Respiratory Syndrome Coronavirus 2 Protein E With Cell Junctions and Polarity PSD-95/Dlg/ZO-1-Containing Proteins.Front Microbiol. 2022 Feb 23;13:829094. doi: 10.3389/fmicb.2022.829094. eCollection 2022. Front Microbiol. 2022. PMID: 35283834 Free PMC article.
-
Allosterism in the PDZ Family.Int J Mol Sci. 2022 Jan 27;23(3):1454. doi: 10.3390/ijms23031454. Int J Mol Sci. 2022. PMID: 35163402 Free PMC article. Review.
-
BCR: a promiscuous fusion partner in hematopoietic disorders.Oncotarget. 2019 Apr 12;10(28):2738-2754. doi: 10.18632/oncotarget.26837. eCollection 2019 Apr 12. Oncotarget. 2019. PMID: 31105873 Free PMC article. Review.
-
Viral PDZ Binding Motifs Influence Cell Behavior Through the Interaction with Cellular Proteins Containing PDZ Domains.Methods Mol Biol. 2021;2256:217-236. doi: 10.1007/978-1-0716-1166-1_13. Methods Mol Biol. 2021. PMID: 34014525 Review.
References
-
- Bezprozvanny I. and Maximov, A. 2001. Classification of PDZ domains. FEBS Lett. 509: 457–462. - PubMed
-
- Bezprozvanny I. and Maximov, A. 2002. PDZ domains: Evolving classification. FEBS Lett. 512: 347–349. - PubMed
-
- Birrane G., Chung, J., and Ladias, J.A. 2003. Novel mode of ligand recognition by the Erbin PDZ domain. J. Biol. Chem. 278: 1399–1402. - PubMed
-
- Brunger A.T., Adams, P.D., Clore, G.M., DeLano, W.L., Gros, P., Grosse-Kunstleve, R.W., Jiang, J.S., Kuszewski, J., Nilges, M., Pannu, N.S., et al. 1998. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54: 905–921. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources

