A fluorescence polarization-based screening assay for nucleic acid polymerase elongation activity
- PMID: 17475199
- PMCID: PMC2713175
- DOI: 10.1016/j.ab.2007.03.039
A fluorescence polarization-based screening assay for nucleic acid polymerase elongation activity
Abstract
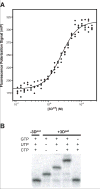
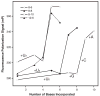
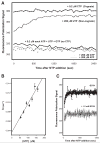
We have devised a simple high-throughput screening compatible fluorescence polarization-based assay that can be used to detect the elongation activity of nucleic acid polymerase enzymes. The assay uses a 5' end-labeled template strand and relies on an increase in the polarization signal from the fluorescent label as it is drawn in toward the active site by the action of the enzyme. If the oligonucleotide is sufficiently short, the fluorescence polarization signal can also be used to detect binding prior to elongation activity. We refer to the nucleic acid substrate as a polymerase elongation template element (PETE) and demonstrate the utility of this PETE assay in a microtiter plate format using the RNA-dependent RNA polymerase from poliovirus to extend a self-priming hairpin RNA. The PETE assay provides an efficient method for screening compounds that may inhibit the nucleic acid binding or elongation activities of polymerases.
Figures

Similar articles
-
A quantitative stopped-flow fluorescence assay for measuring polymerase elongation rates.Anal Biochem. 2009 Aug 1;391(1):45-55. doi: 10.1016/j.ab.2009.04.035. Epub 2009 May 3. Anal Biochem. 2009. PMID: 19406094 Free PMC article.
-
Fluorescence characterization of the transcription bubble in elongation complexes of T7 RNA polymerase.J Mol Biol. 2001 May 4;308(3):465-75. doi: 10.1006/jmbi.2001.4601. J Mol Biol. 2001. PMID: 11327781
-
The T7 RNA polymerase intercalating hairpin is important for promoter opening during initiation but not for RNA displacement or transcription bubble stability during elongation.Biochemistry. 2001 Apr 3;40(13):3882-90. doi: 10.1021/bi002716c. Biochemistry. 2001. PMID: 11300767
-
Fluorescent methods to study transcription initiation and transition into elongation.Exp Suppl. 2014;105:105-30. doi: 10.1007/978-3-0348-0856-9_6. Exp Suppl. 2014. PMID: 25095993 Free PMC article. Review.
-
Structure and function in promoter escape by T7 RNA polymerase.Prog Nucleic Acid Res Mol Biol. 2005;80:323-47. doi: 10.1016/S0079-6603(05)80008-X. Prog Nucleic Acid Res Mol Biol. 2005. PMID: 16164978 Review. No abstract available.
Cited by
-
Fluorescence Polarization-Based Bioassays: New Horizons.Sensors (Basel). 2020 Dec 12;20(24):7132. doi: 10.3390/s20247132. Sensors (Basel). 2020. PMID: 33322750 Free PMC article. Review.
-
In vitro methods for testing antiviral drugs.Biotechnol Adv. 2018 May-Jun;36(3):557-576. doi: 10.1016/j.biotechadv.2017.12.016. Epub 2017 Dec 29. Biotechnol Adv. 2018. PMID: 29292156 Free PMC article. Review.
-
An in vitro fluorescence based study of initiation of RNA synthesis by influenza B polymerase.Nucleic Acids Res. 2017 Apr 7;45(6):3353-3368. doi: 10.1093/nar/gkx043. Nucleic Acids Res. 2017. PMID: 28126917 Free PMC article.
-
A template RNA entry channel in the fingers domain of the poliovirus polymerase.J Mol Biol. 2012 Apr 6;417(4):263-78. doi: 10.1016/j.jmb.2012.01.049. Epub 2012 Feb 1. J Mol Biol. 2012. PMID: 22321798 Free PMC article.
-
High-throughput screening identification of poliovirus RNA-dependent RNA polymerase inhibitors.Antiviral Res. 2011 Sep;91(3):241-51. doi: 10.1016/j.antiviral.2011.06.006. Epub 2011 Jun 21. Antiviral Res. 2011. PMID: 21722674 Free PMC article.
References
-
- Beaulieu PL. Finger loop inhibitors of the HCV NS5B polymerase: discovery and prospects for new HCV therapy. Curr Opin Drug Discov Devel. 2006;9:618–26. - PubMed
-
- Arnold JJ, Cameron CE. Poliovirus RNA-dependent RNA polymerase (3D(pol)). Assembly of stable, elongation-competent complexes by using a symmetrical primer-template substrate (sym/sub) J Biol Chem. 2000;275:5329–36. - PubMed
-
- Ferrer-Orta C, Arias A, Perez-Luque R, Escarmis C, Domingo E, Verdaguer N. Structure of foot-and-mouth disease virus RNA-dependent RNA polymerase and its complex with a template-primer RNA. J Biol Chem. 2004;279:47212–21. - PubMed
-
- Yin YW, Steitz TA. Structural basis for the transition from initiation to elongation transcription in T7 RNA polymerase. Science. 2002;298:1387–95. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

